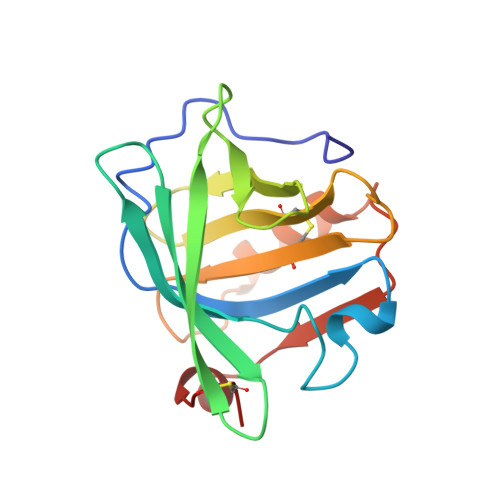
Structural basis of the Tanford transition of bovine beta-lactoglobulin.
Qin, B.Y., Bewley, M.C., Creamer, L.K., Baker, H.M., Baker, E.N., Jameson, G.B.(1998) Biochemistry 37: 14014-14023
- PubMed: 9760236 Search on PubMed
- DOI: https://doi.org/10.1021/bi981016t
- Primary Citation Related Structures:
1BSY, 2BLG, 3BLG - PubMed Abstract:
The structures of the trigonal crystal form of bovine beta-lactoglobulin variant A at pH 6.2, 7.1, and 8.2 have been determined by X-ray diffraction methods at a resolution of 2.56, 2. 24, and 2.49 A, respectively. The corresponding values for R (Rfree) are 0.192 (0.240), 0.234 (0.279), and 0.232 (0.277). The C and N termini as well as two disulfide bonds are clearly defined in these models. The glutamate side chain of residue 89 is buried at pH 6.2 and becomes exposed at pH 7.1 and 8.2. This conformational change, involving the loop 85-90, provides a structural basis for a variety of pH-dependent chemical, physical, and spectroscopic phenomena, collectively known as the Tanford transition.
- Centre for Structural Biology, Institutes of Fundamental Sciences and Molecular Biosciences, Massey University, Palmerston North, New Zealand.
Organizational Affiliation: