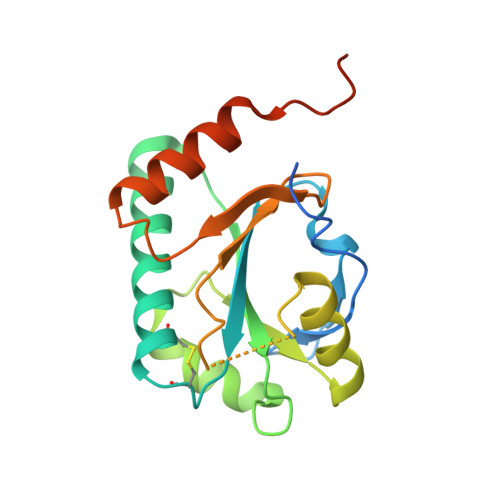
Crystal structure of yeast Sco1.
Abajian, C., Rosenzweig, A.C.(2006) J Biol Inorg Chem 11: 459-466
- PubMed: 16570183 Search on PubMed
- DOI: https://doi.org/10.1007/s00775-006-0096-7
- Primary Citation Related Structures:
2B7J, 2B7K - PubMed Abstract:
The Sco family of proteins are involved in the assembly of the dinuclear CuA site in cytochrome c oxidase (COX), the terminal enzyme in aerobic respiration. These proteins, which are found in both eukaryotes and prokaryotes, are characterized by a conserved CXXXC sequence motif that binds copper ions and that has also been proposed to perform a thiol:disulfide oxidoreductase function. The crystal structures of Saccharomyces cerevisiae apo Sco1 (apo-ySco1) and Sco1 in the presence of copper ions (Cu-ySco1) were determined to 1.8- and 2.3-A resolutions, respectively. Yeast Sco1 exhibits a thioredoxin-like fold, similar to that observed for human Sco1 and a homolog from Bacillus subtilis. The Cu-ySco1 structure, obtained by soaking apo-ySco1 crystals in copper ions, reveals an unexpected copper-binding site involving Cys181 and Cys216, cysteine residues present in ySco1 but not in other homologs. The conserved CXXXC cysteines, Cys148 and Cys152, can undergo redox chemistry in the crystal. An essential histidine residue, His239, is located on a highly flexible loop, denoted the Sco loop, and can adopt positions proximal to both pairs of cysteines. Interactions between ySco1 and its partner proteins yeast Cox17 and yeast COX2 are likely to occur via complementary electrostatic surfaces. This high-resolution model of a eukaryotic Sco protein provides new insight into Sco copper binding and function.
- Department of Biochemistry, Molecular Biology, and Cell Biology, Northwestern University, Evanston, IL 60208, USA.
Organizational Affiliation: