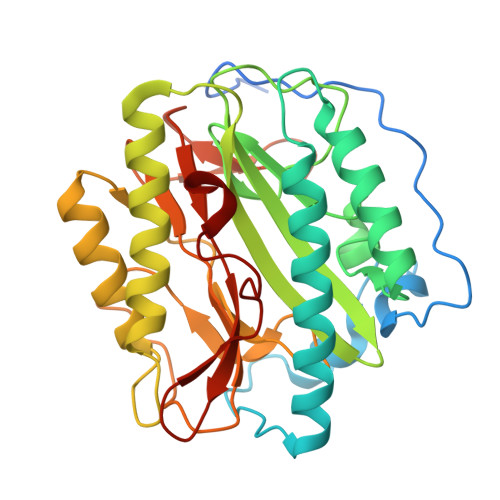
Structural Basis for the Functional Differences between Type I and Type II Human Methionine Aminopeptidases(,).
Addlagatta, A., Hu, X., Liu, J.O., Matthews, B.W.(2005) Biochemistry 44: 14741-14749
- PubMed: 16274222
- DOI: https://doi.org/10.1021/bi051691k
- Primary Citation Related Structures:
2B3H, 2B3K, 2B3L - PubMed Abstract:
Determination of the crystal structure of human MetAP1 makes it possible, for the first time, to compare the structures of a Type I and a Type II methionine aminopeptidase (MetAP) from the same organism. Comparison of the Type I enzyme with the previously reported complex of ovalicin with Type II MetAP shows that the active site of the former is reduced in size and would incur steric clashes with the bound inhibitor. This explains why ovalicin and related anti-angiogenesis inhibitors target Type II human MetAP but not Type I. The differences in both size and shape of the active sites between MetAP1 and MetAP2 also help to explain their different substrate specificity. In the presence of excess Co(2+), a third cobalt ion binds in the active site region, explaining why metal ions in excess can be inhibitory. Also, the N-terminal region of the protein contains three distinct Pro-x-x-Pro motifs, supporting the prior suggestion that this region of the protein may participate in binding to the ribosome.
- Institute of Molecular Biology, Howard Hughes Medical Institute and Department of Physics, 1229 University of Oregon, Eugene, Oregon 97403-1229, USA.
Organizational Affiliation: