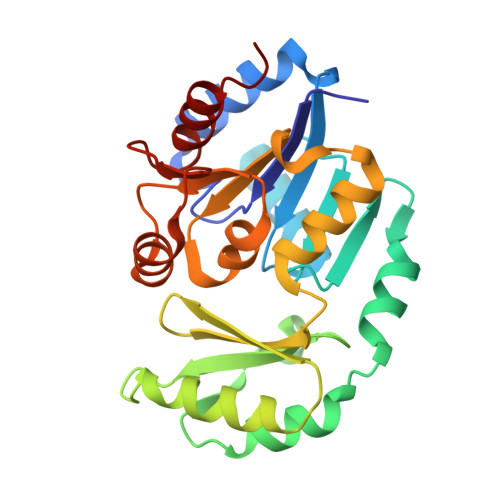
Crystal structure of a cyanobacterial sucrose-phosphatase in complex with glucose-containing disaccharides
Fieulaine, S., Lunn, J.E., Ferrer, J.-L.(2007) Proteins 68: 796-801
- PubMed: 17510968 Search on PubMed
- DOI: https://doi.org/10.1002/prot.21481
- Primary Citation Related Structures:
2B1Q, 2B1R, 2D2V - Institut de Biologie Structurale J-P. Ebel, CEA/CNRS/UJF, Grenoble, France.
Organizational Affiliation: