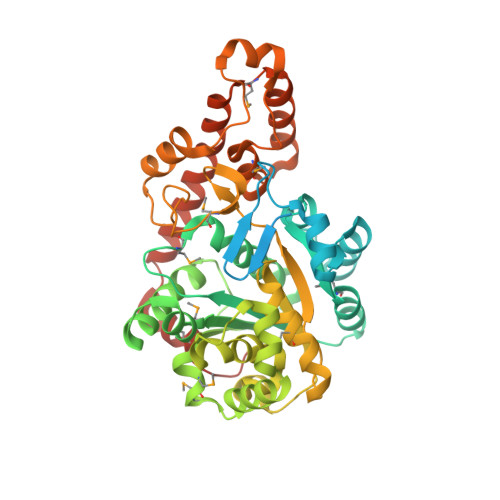
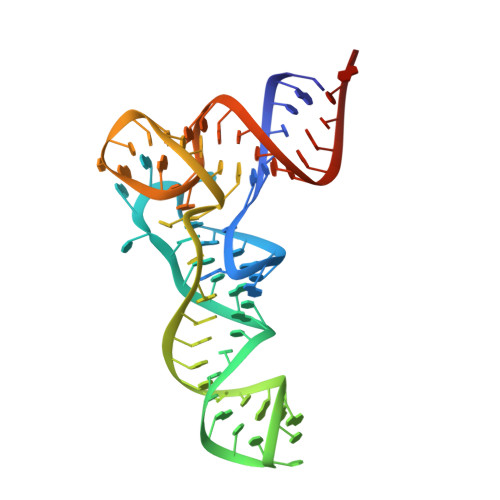
Two conformations of a crystalline human tRNA synthetase-tRNA complex: implications for protein synthesis.
Yang, X.L., Otero, F.J., Ewalt, K.L., Liu, J., Swairjo, M.A., Kohrer, C., RajBhandary, U.L., Skene, R.J., McRee, D.E., Schimmel, P.(2006) EMBO J 25: 2919-2929
- PubMed: 16724112
- DOI: https://doi.org/10.1038/sj.emboj.7601154
- Primary Citation Related Structures:
2AZX - PubMed Abstract:
Aminoacylation of tRNA is the first step of protein synthesis. Here, we report the co-crystal structure of human tryptophanyl-tRNA synthetase and tRNATrp. This enzyme is reported to interact directly with elongation factor 1alpha, which carries charged tRNA to the ribosome. Crystals were generated from a 50/50% mixture of charged and uncharged tRNATrp. These crystals captured two conformations of the complex, which are nearly identical with respect to the protein and a bound tryptophan. They are distinguished by the way tRNA is bound. In one, uncharged tRNA is bound across the dimer, with anticodon and acceptor stem interacting with separate subunits. In this cross-dimer tRNA complex, the class I enzyme has a class II-like tRNA binding mode. This structure accounts for biochemical investigations of human TrpRS, including species-specific charging. In the other conformation, presumptive aminoacylated tRNA is bound only by the anticodon, the acceptor stem being free and having space to interact precisely with EF-1alpha, suggesting that the product of aminoacylation can be directly handed off to EF-1alpha for the next step of protein synthesis.
- Department of Molecular Biology, The Scripps Research Institute, La Jolla, CA 92037, USA. xlyang@scripps.edu
Organizational Affiliation: