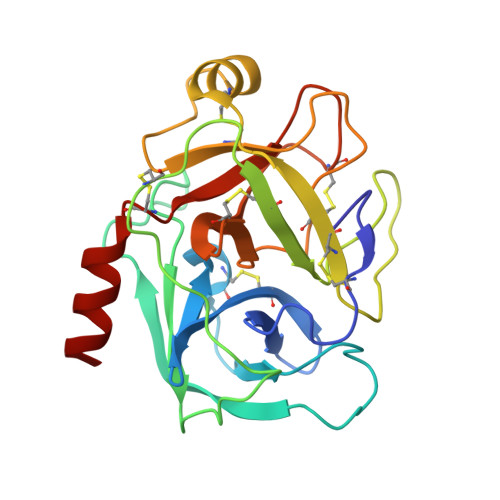
Structure of the complex of trypsin with a highly potent synthetic inhibitor at 0.97 A resolution.
Sherawat, M., Kaur, P., Perbandt, M., Betzel, C., Slusarchyk, W.A., Bisacchi, G.S., Chang, C., Jacobson, B.L., Einspahr, H.M., Singh, T.P.(2007) Acta Crystallogr D Biol Crystallogr 63: 500-507
- PubMed: 17372355
- DOI: https://doi.org/10.1107/S090744490700697X
- Primary Citation Related Structures:
2AYW - PubMed Abstract:
The structure of the complex formed between bovine beta-trypsin and the highly potent synthetic inhibitor 2-{3'-[5'-methoxy-2'-(N-p-diaminomethylphenyl)amido]-1'-pyrido}-5-(N-2''-t-butylethanol)amidobenzoic acid (C(28)H(32)N(5)O(6)) has been determined at 0.97 A resolution. X-ray intensity data were collected to 0.97 A under cryocooled conditions. The structure was refined anisotropically using REFMAC5 and SHELX-97 to R(cryst) factors of 13.4 and 12.6% and R(free) factors of 15.7 and 16.3%, respectively. Several regions of the main chain and side chains that have not been previously observed were clearly defined in the present structure. H atoms are indicated as significant peaks in an |F(o) - F(c)| difference map, which accounts for an estimated 35% of all H atoms at the 2.5sigma level. The C, N and O atoms are definitively differentiated in the electron-density maps. The amido part of the inhibitor occupies the specificity pocket and the remainder fills the remaining part of the ligand-binding cleft and interacts with the enzyme through an extensive network of hydrogen bonds. The inhibitor distorts the stereochemistry of the catalytic triad, Ser195-His57-Asp102, thereby blocking the proton-relay process of the active site by preventing the formation of the crucial hydrogen bond between His57 N(delta1) and Asp102 O(delta1).
- Department of Biophysics, All India Institute of Medical Sciences, New Delhi 110029, India.
Organizational Affiliation: