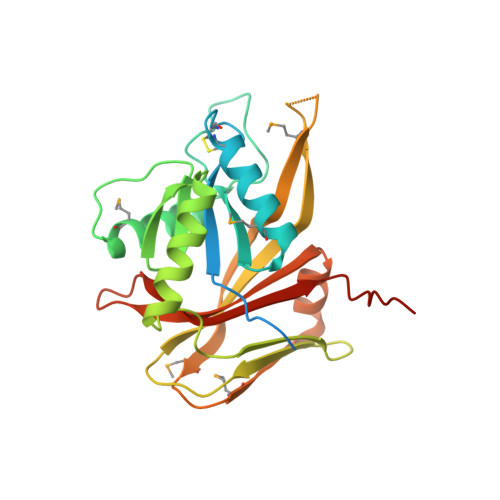
Crystal Structure of the Hypothetical Protein AGR_C_4864 from Agrobacterium tumefaciens, NESG target AtR35
Forouhar, F., Abashidze, M., Benach, J., Xiao, R., Janjua, H., Conover, K., Acton, T.B., Montelione, G.T., Tong, L., Hunt, J.F.To be published.