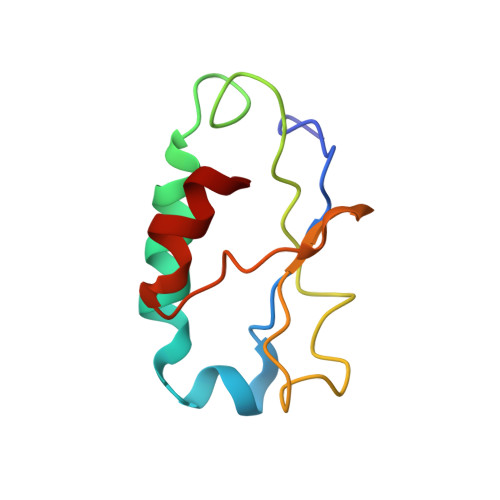
Solution structure of HndAc: a thioredoxin-like domain involved in the NADP-reducing hydrogenase complex
Nouailler, M., Morelli, X., Bornet, O., Chetrit, B., Dermoun, Z., Guerlesquin, F.(2006) Protein Sci 15: 1369-1378
- PubMed: 16731971 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.051916606
- Primary Citation Related Structures:
2AUV - PubMed Abstract:
The NADP-reducing hydrogenase complex from Desulfovibrio fructosovorans is a heterotetramer encoded by the hndABCD operon. Sequence analysis indicates that the HndC subunit (52 kDa) corresponds to the NADP-reducing unit, and the HndD subunit (63.5 kDa) is homologous to Clostridium pasteurianum hydrogenase. The role of HndA and HndB subunits (18.8 kDa and 13.8 kDa, respectively) in the complex remains unknown. The HndA subunit belongs to the [2Fe-2S] ferredoxin family typified by C. pasteurianum ferredoxin. HndA is organized into two independent structural domains, and we report in the present work the NMR structure of its C-terminal domain, HndAc. HndAc has a thioredoxin-like fold consisting in four beta-strands and two relatively long helices. The [2Fe-2S] cluster is located near the surface of the protein and bound to four cysteine residues particularly well conserved in this class of proteins. Electron exchange between the HndD N-terminal [2Fe-2S] domain (HndDN) and HndAc has been previously evidenced, and in the present studies we have mapped the binding site of the HndDN domain on HndAc. A structural analysis of HndB indicates that it is a FeS subunit with 41% similarity with HndAc and it contains a possible thioredoxin-like fold. Our data let us propose that HndAc and HndB can form a heterodimeric intermediate in the electron transfer between the hydrogenase (HndD) active site and the NADP reduction site in HndC.
- Unité de Bioénergétique et Ingénierie des Protéines, IBSM-CNRS, Marseille Cedex 20, France.
Organizational Affiliation: