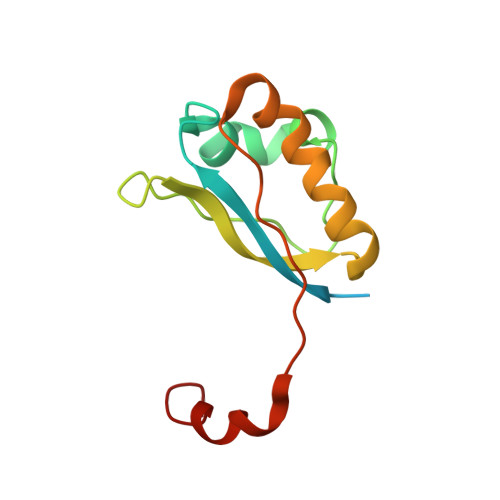
Solution structure of the Escherichia coli protein ydhR: A putative mono-oxygenase.
Revington, M., Semesi, A., Yee, A., Shaw, G.S.(2005) Protein Sci 14: 3115-3120
- PubMed: 16260765
- DOI: https://doi.org/10.1110/ps.051809305
- Primary Citation Related Structures:
2ASY - PubMed Abstract:
YdhR is a 101-residue conserved protein from Escherichia coli. Sequence searches reveal that the protein has >50% identity to proteins found in a variety of other bacterial genomes. Using size exclusion chromatography and fluorescence spectroscopy, we determined that ydhR exists in a dimeric state with a dissociation constant of approximately 40 nM. The three-dimensional structure of dimeric ydhR was determined using NMR spectroscopy. A total of 3400 unambiguous NOEs, both manually and automatically assigned, were used for the structure calculation that was refined using an explicit hydration shell. A family of 20 structures was obtained with a backbone RMSD of 0.48 A for elements of secondary structure. The structure reveals a dimeric alpha,beta fold characteristic of the alpha+beta barrel superfamily of proteins. Bioinformatic approaches were used to show that ydhR likely belongs to a recently identified group of mono-oxygenase proteins that includes ActVA-Orf6 and YgiN and are involved in the oxygenation of polyaromatic ring compounds.
- Department of Biochemistry, The University of Western Ontario, London, ON, N6A 5C1, Canada.
Organizational Affiliation: