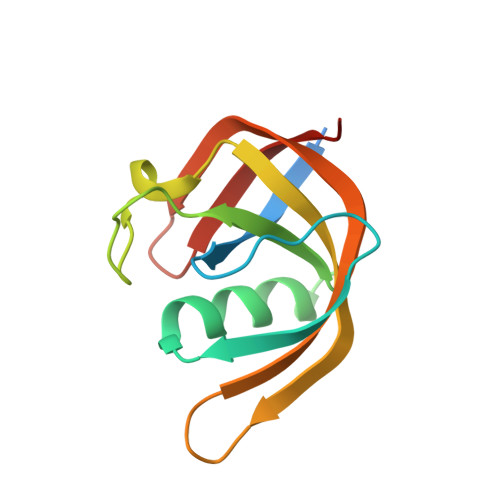
Crystal structure of the N-terminal domain of E. coli Lon protease.
Li, M., Rasulova, F., Melnikov, E.E., Rotanova, T.V., Gustchina, A., Maurizi, M.R., Wlodawer, A.(2005) Protein Sci 14: 2895-2900
- PubMed: 16199667
- DOI: https://doi.org/10.1110/ps.051736805
- Primary Citation Related Structures:
2ANE - PubMed Abstract:
We report here the first crystal structure of the N-terminal domain of an A-type Lon protease. Lon proteases are ubiquitous, multidomain, ATP-dependent enzymes with both highly specific and non-specific protein binding, unfolding, and degrading activities. We expressed and purified a stable, monomeric 119-amino acid N-terminal subdomain of the Escherichia coli A-type Lon protease and determined its crystal structure at 2.03 A (Protein Data Bank [PDB] code 2ANE). The structure was solved in two crystal forms, yielding 14 independent views. The domain exhibits a unique fold consisting primarily of three twisted beta-sheets and a single long alpha-helix. Analysis of recent PDB depositions identified a similar fold in BPP1347 (PDB code 1ZBO), a 203-amino acid protein of unknown function from Bordetella parapertussis, crystallized as part of a structural genomics effort. BPP1347 shares sequence homology with Lon N-domains and with a family of other independently expressed proteins of unknown functions. We postulate that, as is the case in Lon proteases, this structural domain represents a general protein and polypeptide interaction domain.
- Macromolecular Crystallography Laboratory, National Cancer Institute, Building 536, Room 5, Frederick, MD 21702-1201, USA.
Organizational Affiliation: