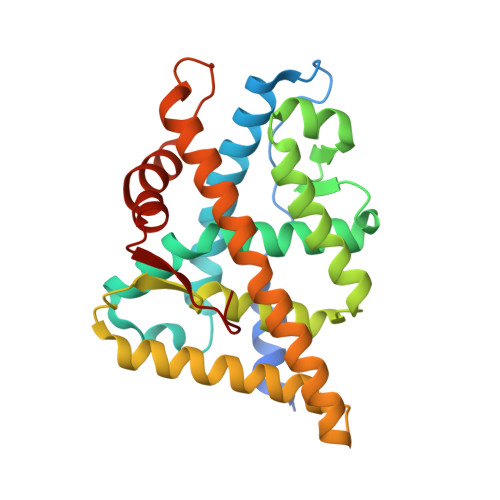
Comparison of crystal structures of human androgen receptor ligand-binding domain complexed with various agonists reveals molecular determinants responsible for binding affinity.
Pereira de Jesus-Tran, K., Cote, P.-L., Cantin, L., Blanchet, J., Labrie, F., Breton, R.(2006) Protein Sci 15: 987-999
- PubMed: 16641486 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.051905906
- Primary Citation Related Structures:
2AM9, 2AMA, 2AMB - PubMed Abstract:
Androgens exert their effects by binding to the highly specific androgen receptor (AR). In addition to natural potent androgens, AR binds a variety of synthetic agonist or antagonist molecules with different affinities. To identify molecular determinants responsible for this selectivity, we have determined the crystal structure of the human androgen receptor ligand-binding domain (hARLBD) in complex with two natural androgens, testosterone (Testo) and dihydrotestosterone (DHT), and with an androgenic steroid used in sport doping, tetrahydrogestrinone (THG), at 1.64, 1.90, and 1.75 A resolution, respectively. Comparison of these structures first highlights the flexibility of several residues buried in the ligand-binding pocket that can accommodate a variety of ligand structures. As expected, the ligand structure itself (dimension, presence, and position of unsaturated bonds that influence the geometry of the steroidal nucleus or the electronic properties of the neighboring atoms, etc.) determines the number of interactions it can make with the hARLBD. Indeed, THG--which possesses the highest affinity--establishes more van der Waals contacts with the receptor than the other steroids, whereas the geometry of the atoms forming electrostatic interactions at both extremities of the steroid nucleus seems mainly responsible for the higher affinity measured experimentally for DHT over Testo. Moreover, estimation of the ligand-receptor interaction energy through modeling confirms that even minor modifications in ligand structure have a great impact on the strength of these interactions. Our crystallographic data combined with those obtained by modeling will be helpful in the design of novel molecules with stronger affinity for the AR.
- Oncology and Molecular Endocrinology Research Center, Laval University Medical Center (CHUL) and Laval University, Québec, QC G1V 4G2, Canada.
Organizational Affiliation: