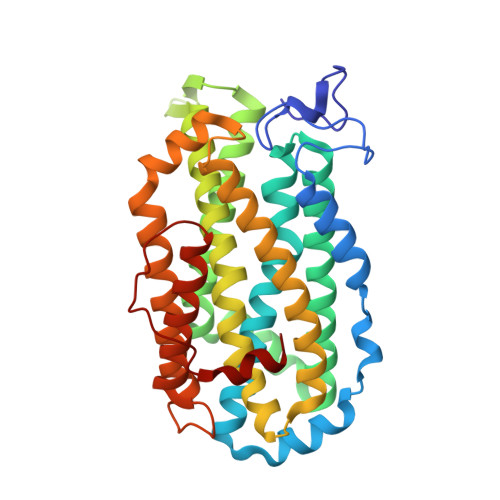
Structure of Escherichia coli ribonucleotide reductase R2 in space group P6122.
Sommerhalter, M., Saleh, L., Bollinger, J.M., Rosenzweig, A.C.(2005) Acta Crystallogr D Biol Crystallogr 61: 1649-1654
- PubMed: 16301799 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905034062
- Primary Citation Related Structures:
2ALX - PubMed Abstract:
A new crystal form of wild-type ribonucleotide reductase R2 from Escherichia coli was obtained. Crystals grow in space group P6(1)22 with one R2 monomer in the asymmetric unit. A twofold crystallographic symmetry axis generates the physiological dimeric form of R2. Co-crystallization with CoCl(2) or MnCl(2) results in full occupancy of the dinuclear metal site. The structure of the Mn(II)-loaded form was determined to 2.6 Angstroms resolution by molecular replacement. The crystallization conditions, backbone conformation, crystal-packing interactions and metal centers are compared with those of previously determined crystal forms.
- Department of Biochemistry, Northwestern University, Evanston, IL 60208, USA.
Organizational Affiliation: