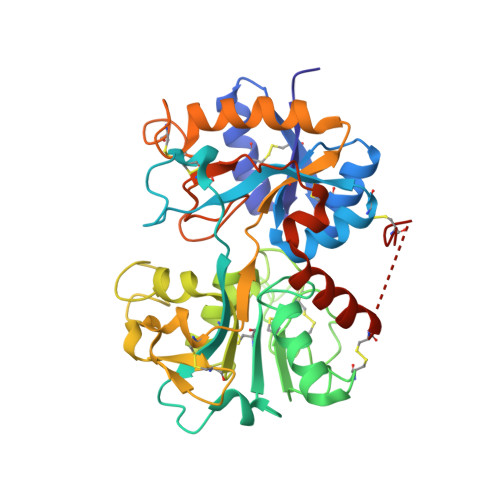
Detection of new binding site in the C-terminal lobe of lactoferrin:Crystal structure of the complex formed between bovine lactoferrin and a tetrasaccharide at 2.1A resolution
Singh, N., Jabeen, T., Sharma, S., Bhushan, A., Singh, T.P.To be published.