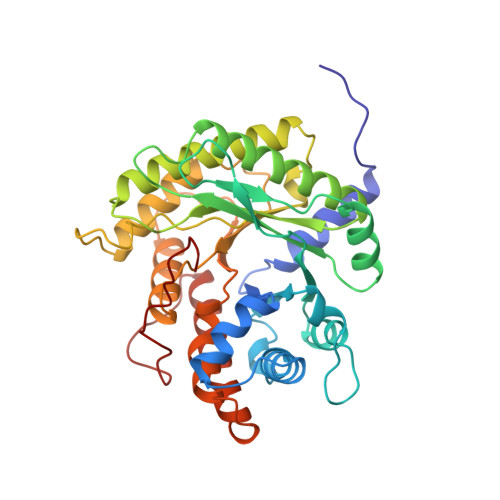
Crystal structure of human muscle aldolase complexed with fructose 1,6-bisphosphate: mechanistic implications.
Dalby, A., Dauter, Z., Littlechild, J.A.(1999) Protein Sci 8: 291-297
- PubMed: 10048322 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1110/ps.8.2.291
- Primary Citation Related Structures:
2ALD, 4ALD - PubMed Abstract:
Fructose 1,6-bisphosphate aldolase catalyzes the reversible cleavage of fructose 1,6-bisphosphate and fructose 1-phosphate to dihydroxyacetone phosphate and either glyceraldehyde 3-phosphate or glyceraldehyde, respectively. Catalysis involves the formation of a Schiff's base intermediate formed at the epsilon-amino group of Lys229. The existing apo-enzyme structure was refined using the crystallographic free-R-factor and maximum likelihood methods that have been shown to give improved structural results that are less subject to model bias. Crystals were also soaked with the natural substrate (fructose 1,6-bisphosphate), and the crystal structure of this complex has been determined to 2.8 A. The apo structure differs from the previous Brookhaven-deposited structure (1ald) in the flexible C-terminal region. This is also the region where the native and complex structures exhibit differences. The conformational changes between native and complex structure are not large, but the observed complex does not involve the full formation of the Schiff's base intermediate, and suggests a preliminary hydrogen-bonded Michaelis complex before the formation of the covalent complex.
- Department of Chemistry and Biological Sciences, Exeter University, United Kingdom.
Organizational Affiliation: