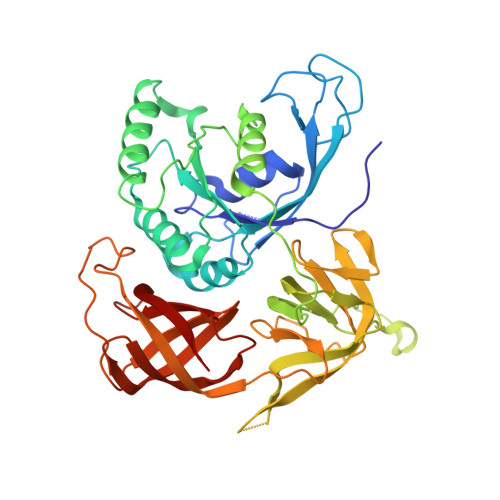
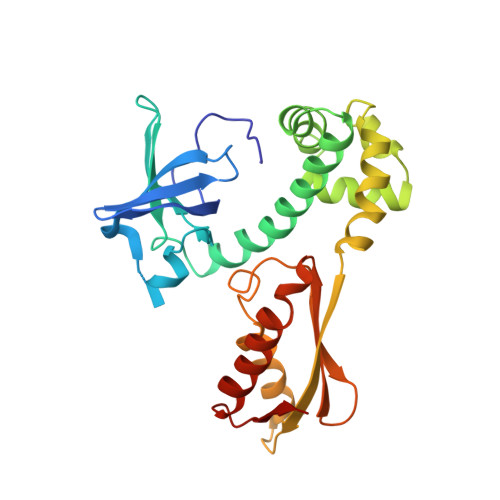
Structural Switch of the gamma Subunit in an Archaeal aIF2alphagamma Heterodimer
Yatime, L., Mechulam, Y., Blanquet, S., Schmitt, E.(2006) Structure 14: 119-128
- PubMed: 16407071
- DOI: https://doi.org/10.1016/j.str.2005.09.020
- Primary Citation Related Structures:
2AHO - PubMed Abstract:
Eukaryotic and archaeal initiation factors 2 (e/aIF2) are heterotrimeric proteins (alphabetagamma) supplying the small subunit of the ribosome with methionylated initiator tRNA. This study reports the crystallographic structure of an aIF2alphagamma heterodimer from Sulfolobus solfataricus bound to Gpp(NH)p-Mg(2+). aIF2gamma is in a closed conformation with the G domain packed on domains II and III. The C-terminal domain of aIF2alpha interacts with domain II of aIF2gamma. Conformations of the two switch regions involved in GTP binding are similar to those encountered in an EF1A:GTP:Phe-tRNA(Phe) complex. Comparison with the EF1A structure suggests that only the gamma subunit of the aIF2alphagamma heterodimer contacts tRNA. Because the alpha subunit markedly reinforces the affinity of tRNA for the gamma subunit, a contribution of the alpha subunit to the switch movements observed in the gamma structure is considered.
- Laboratoire de Biochimie, Unité Mixte de Recherche 7654, CNRS-Ecole Polytechnique, F-91128 Palaiseau cedex, France.
Organizational Affiliation: