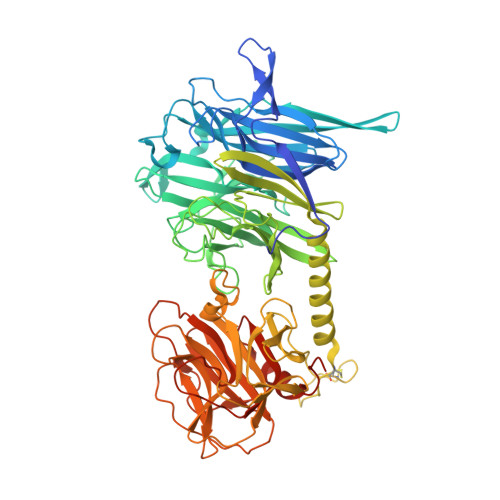
Structural and Kinetic Analysis of Two Covalent Sialosyl-Enzyme Intermediates on Trypanosoma rangeli Sialidase.
Watts, A.G., Oppezzo, P., Withers, S.G., Alzari, P.M., Buschiazzo, A.(2006) J Biological Chem 281: 4149-4155
- PubMed: 16298994 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M510677200
- Primary Citation Related Structures:
2A75, 2AGS, 2FHR - PubMed Abstract:
Trypanosoma rangeli sialidase is a glycoside hydrolase (family GH33) that catalyzes the cleavage of alpha-2-->3-linked sialic acid residues from sialoglycoconjugates with overall retention of anomeric configuration. Retaining glycosidases usually operate through a ping-pong mechanism, wherein a covalent intermediate is formed between the carbohydrate and an active site carboxylic acid of the enzyme. Sialidases, instead, appear to use a tyrosine as the catalytic nucleophile, leaving the possibility of an essentially different catalytic mechanism. Indeed, a direct nucleophilic role for a tyrosine was shown for the homologous trans-sialidase from Trypanosoma cruzi, although itself not a typical sialidase. Here we present the three-dimensional structures of the covalent glycosyl-enzyme complexes formed by the T. rangeli sialidase with two different mechanism-based inactivators at 1.9 and 1.7 Angstroms resolution. To our knowledge, these are the first reported structures of enzymatically competent covalent intermediates for a strictly hydrolytic sialidase. Kinetic analyses have been carried out on the formation and turnover of both intermediates, showing that structural modifications to these inactivators can be used to modify the lifetimes of covalent intermediates. These results provide further evidence that all sialidases likely operate through a similar mechanism involving the transient formation of a covalently sialylated enzyme. Furthermore, we believe that the ability to "tune" the inactivation and reactivation rates of mechanism-based inactivators toward specific enzymes represents an important step toward developing this class of inactivators into therapeutically useful compounds.
- Department of Chemistry, University of British Columbia, Vancouver, Canada.
Organizational Affiliation: