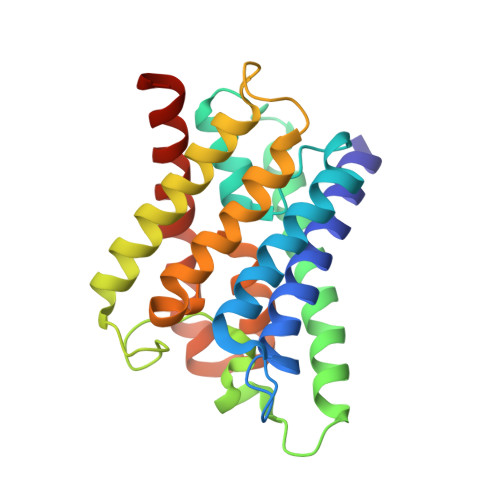
Crystal Structure of AqpZ Tetramer Reveals Two Distinct Arg-189 Conformations Associated with Water Permeation through the Narrowest Constriction of the Water-conducting Channel.
Jiang, J., Daniels, B.V., Fu, D.(2006) J Biological Chem 281: 454-460
- PubMed: 16239219 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M508926200
- Primary Citation Related Structures:
2ABM - PubMed Abstract:
AqpZ is a homotetramer of four water-conducting channels that facilitate rapid water movements across the plasma membrane of Escherichia coli. Here we report a 3.2 angstroms crystal structure of the tetrameric AqpZ (tAqpZ). All channel-lining residues in the four monomeric channels are found orientated in nearly identical positions with one marked exception at the narrowest channel constriction, where the side chain of a highly conserved Arg-189 adopts two distinct conformational orientations. In one of the four monomers, the guanidino group of Arg-189 points toward the periplasmic vestibule, opening up the constriction to accommodate the binding of a water molecule through a tridentate H-bond. In the other three monomers, the Arg-189 guanidino group bends over to form an H-bond with carbonyl oxygen of the Thr-183, thus occluding the channel. Therefore, the tAqpZ structure reveals two distinct Arg-189 confirmations associated with water permeation through the channel constrictions. Alternation between the two Arg-189 conformations disrupts continuous flow of water, thus regulating the open probability of the water pore. Further, the difference in Arg-189 displacements is correlated with a strong electron density found between the first transmembrane helices of two open channels, suggesting that the observed Arg-189 conformations are stabilized by asymmetrical subunit interactions in tAqpZ.
- Department of Biology, Brookhaven National Laboratory, Upton, New York 11973, USA.
Organizational Affiliation: