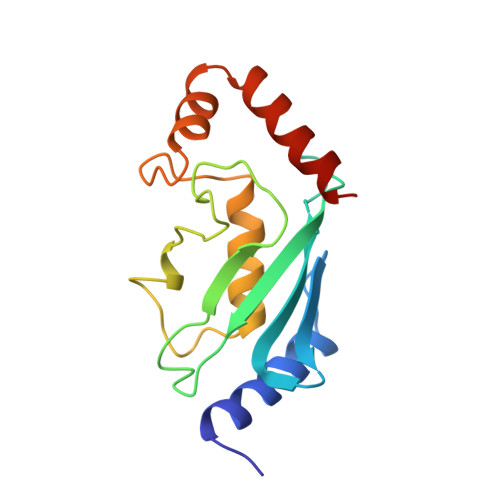
Three-dimensional structure of a ubiquitin-conjugating enzyme (E2).
Cook, W.J., Jeffrey, L.C., Sullivan, M.L., Vierstra, R.D.(1992) J Biological Chem 267: 15116-15121
- PubMed: 1321826 Search on PubMed
- DOI: https://doi.org/10.2210/pdb1aak/pdb
- Primary Citation Related Structures:
2AAK - PubMed Abstract:
The x-ray crystal structure of a recombinant ubiquitin-conjugating enzyme (E2) encoded by the UBC1 gene of the plant Arabidopsis thaliana has been determined with the use of multiple isomorphous replacement techniques and refined at 2.4-A resolution by simulated annealing and restrained least-squares. This E2 is an alpha/beta protein, with four alpha-helices and a four-stranded antiparallel beta-sheet. The NH2 and COOH termini, which may be important for interaction with other enzymes and substrates in the ubiquitin-conjugation pathway, are on the opposite side of the molecule from the cysteine residue that binds to the COOH terminus of ubiquitin. This structure should now allow for the rational analysis of E2 function by in vitro mutagenesis and facilitate the effective design of E2s with unique specificities or catalytic functions.
- Department of Pathology, University of Alabama, Birmingham 35294.
Organizational Affiliation: