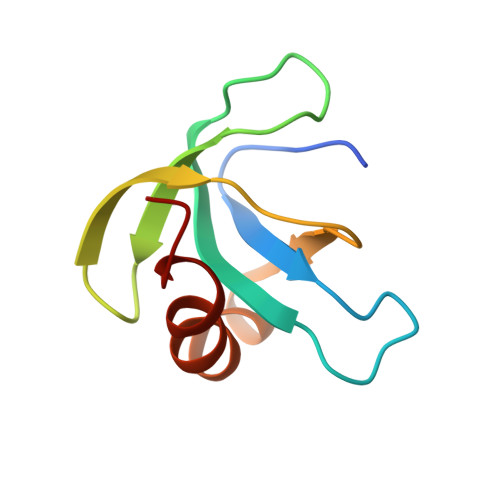
Solution structure of the conserved hypothetical protein Rv2302 from Mycobacterium tuberculosis.
Buchko, G.W., Kim, C.Y., Terwilliger, T.C., Kennedy, M.A.(2006) J Bacteriol 188: 5993-6001
- PubMed: 16885468 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1128/JB.00460-06
- Primary Citation Related Structures:
2A7Y - PubMed Abstract:
The Mycobacterium tuberculosis protein Rv2302 (80 residues; molecular mass of 8.6 kDa) has been characterized using nuclear magnetic resonance (NMR) and circular dichroism (CD) spectroscopy. While the biochemical function of Rv2302 is still unknown, recent microarray analyses show that Rv2302 is upregulated in response to starvation and overexpression of heat shock proteins and, consequently, may play a role in the biochemical processes associated with these events. Rv2302 is a monomer in solution as shown by size exclusion chromatography and NMR spectroscopy. CD spectroscopy suggests that Rv2302 partially unfolds upon heating and that this unfolding is reversible. Using NMR-based methods, the solution structure of Rv2302 was determined. The protein contains a five-strand, antiparallel beta-sheet core with one C-terminal alpha-helix (A61 to A75) nestled against its side. Hydrophobic interactions between residues in the alpha-helix and beta-strands 3 and 4 hold the alpha-helix near the beta-sheet core. The electrostatic potential on the solvent-accessible surface is primarily negative with the exception of a positive arginine pocket composed of residues R18, R70, and R74. Steady-state {(1)H}-(15)N heteronuclear nuclear Overhauser effects indicate that the protein's core is rigid on the picosecond timescale. The absence of amide cross-peaks for residues G13 to H19 in the (1)H-(15)N heteronuclear single quantum correlation spectrum suggests that this region, a loop between beta-strands 1 and 2, undergoes motion on the millisecond to microsecond timescale. Dali searches using the structure closest to the average structure do not identify any high similarities to any other known protein structure, suggesting that the structure of Rv2302 may represent a novel protein fold.
- Fundamental Sciences, Biological Sciences Division, Battelle, Pacific Northwest National Laboratory, P.O. Box 999, Mail Stop K8-98, Richland, WA 99352, USA.
Organizational Affiliation: