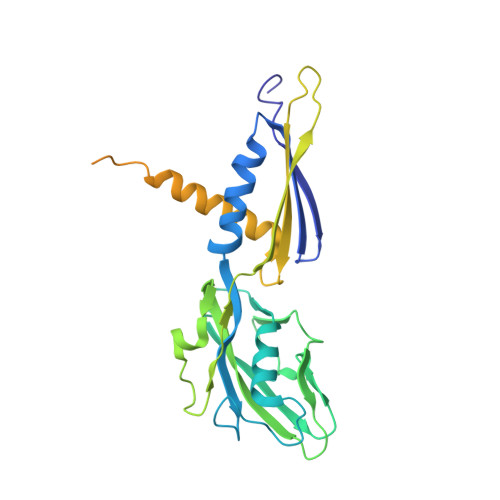
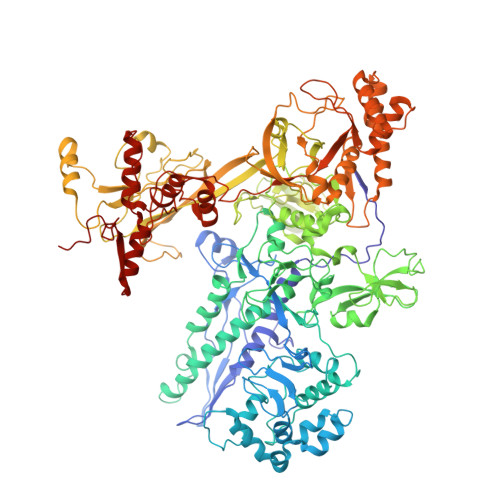
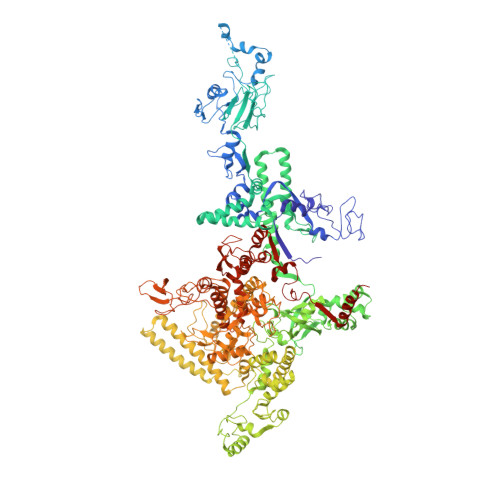
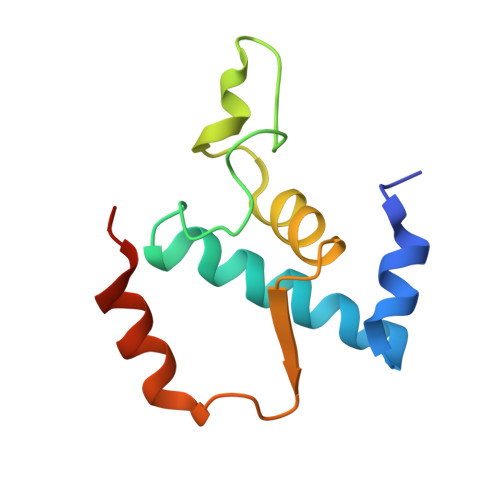
Allosteric modulation of the RNA polymerase catalytic reaction is an essential component of transcription control by rifamycins.
Artsimovitch, I., Vassylyeva, M.N., Svetlov, D., Svetlov, V., Perederina, A., Igarashi, N., Matsugaki, N., Wakatsuki, S., Tahirov, T.H., Vassylyev, D.G.(2005) Cell 122: 351-363
- PubMed: 16096056 Search on PubMed
- DOI: https://doi.org/10.1016/j.cell.2005.07.014
- Primary Citation Related Structures:
2A68, 2A69, 2A6E - PubMed Abstract:
Rifamycins, the clinically important antibiotics, target bacterial RNA polymerase (RNAP). A proposed mechanism in which rifamycins sterically block the extension of nascent RNA beyond three nucleotides does not alone explain why certain RNAP mutations confer resistance to some but not other rifamycins. Here we show that unlike rifampicin and rifapentin, and contradictory to the steric model, rifabutin inhibits formation of the first and second phosphodiester bonds. We report 2.5 A resolution structures of rifabutin and rifapentin complexed with the Thermus thermophilus RNAP holoenzyme. The structures reveal functionally important distinct interactions of antibiotics with the initiation sigma factor. Strikingly, both complexes lack the catalytic Mg2+ ion observed in the apo-holoenzyme, whereas an increase in Mg2+ concentration confers resistance to rifamycins. We propose that a rifamycin-induced signal is transmitted over approximately 19 A to the RNAP active site to slow down catalysis. Based on structural predictions, we designed enzyme substitutions that apparently interrupt this allosteric signal.
- Department of Microbiology, The Ohio State University, 484 West 12th Avenue, Columbus, Ohio 43210, USA.
Organizational Affiliation: