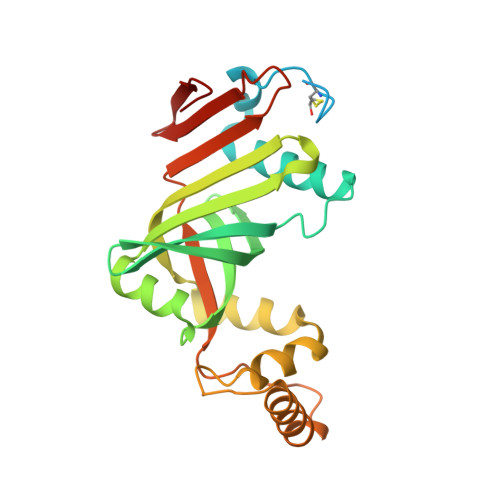
Crystal structure of a putative pyridoxine 5'-phosphate oxidase (Rv2607) from Mycobacterium tuberculosis.
Pedelacq, J.-D., Rho, B.-S., Kim, C.-Y., Waldo, G.S., Lekin, T.P., Segelke, B.W., Rupp, B., Hung, L.-W., Kim, S.-I., Terwilliger, T.C.(2005) Proteins 62: 563-569
- PubMed: 16374842 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20824
- Primary Citation Related Structures:
2A2J - PubMed Abstract:
The three-dimensional structure of Rv2607, a putative pyridoxine 5'-phosphate oxidase (PNPOx) from Mycobacterium tuberculosis, has been determined by X-ray crystallography to 2.5 A resolution. Rv2607 has a core domain similar to known PNPOx structures with a flavin mononucleotide (FMN) cofactor. Electron density for two FMN at the dimer interface is weak despite the bright yellow color of the protein solution and crystal. The shape and size of the putative binding pocket is markedly different from that of members of the PNPOx family, which may indicate some significant changes in the FMN binding mode of this protein relative to members of the family.
- Bioscience Division, MS M888, Los Alamos National Laboratory, Los Alamos, New Mexico 87545, USA. jpdlcq@lanl.gov
Organizational Affiliation: