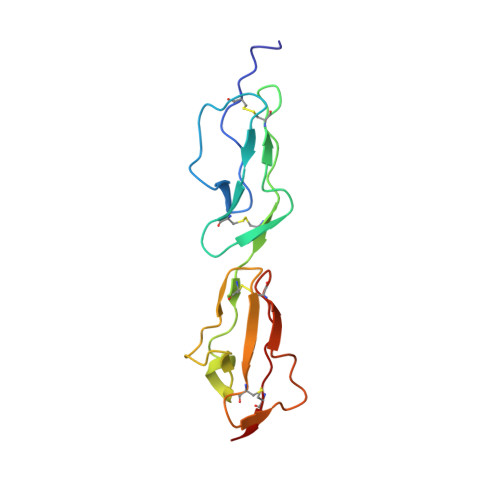
Disease-Associated Sequence Variations Congregate in a Polyanion Recognition Patch on Human Factor H Revealed in Three-Dimensional Structure.
Herbert, A.P., Uhrin, D., Lyon, M., Pangburn, M.K., Barlow, P.N.(2006) J Biological Chem 281: 16512
- PubMed: 16533809 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M513611200
- Primary Citation Related Structures:
2BZM - PubMed Abstract:
Mutations and polymorphisms in the regulator of complement activation, factor H, have been linked to atypical hemolytic uremic syndrome (aHUS), membranoproliferative glomerulonephritis, and age-related macular degeneration. Many aHUS patients carry mutations in the two C-terminal modules of factor H, which normally confer upon this abundant 155-kDa plasma glycoprotein its ability to selectively bind self-surfaces and prevent them from inappropriately triggering the complement cascade via the alternative pathway. In the current study, the three-dimensional solution structure of the C-terminal module pair of factor H has been determined. A binding site for a fully sulfated heparin-derived tetrasaccharide has been delineated using chemical shift mapping and the C3d/C3b-binding site inferred from sequence comparisons and computational docking. The resultant information allows assessment of the likely consequences of aHUS-associated amino acid substitutions in this critical region of factor H. It is striking that, excepting those likely to perturb the three-dimensional structure, aHUS-associated missense mutations congregate in the polyanion-binding site delineated in this study, thus potentially disrupting a vital mechanism for control of complement on self-surfaces in the microvasculature of the kidney. It is intriguing that a single nucleotide polymorphism predisposing to age-related macular degeneration occupies another region of factor H that harbors a polyanion-binding site.
- Edinburgh Biomolecular NMR Unit, University of Edinburgh, West mains Road, Edinburgh EH9 3JJ, United Kingdom.
Organizational Affiliation: