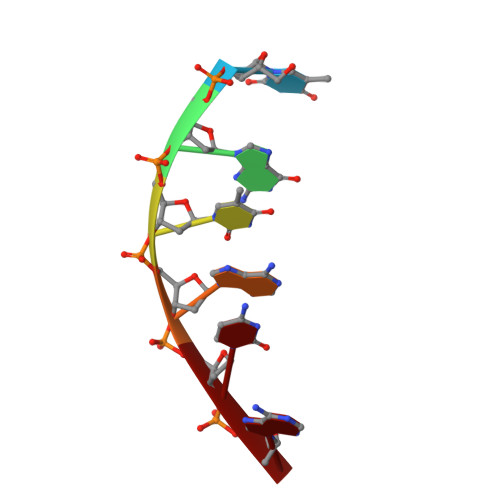
Factors affecting DNA sequence selectivity of nogalamycin intercalation: the crystal structure of d(TGTACA)2-nogalamycin2.
Smith, C.K., Brannigan, J.A., Moore, M.H.(1996) J Mol Biology 263: 237-258
- PubMed: 8913304 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1996.0572
- Primary Citation Related Structures:
258D - PubMed Abstract:
As part of an investigation into the sequence selectivity of the nogalamycin-DNA interaction, the 1.58 A structure of nogalamycin complexed with d5'(TGTACA)2 has been determined by single-crystal X-ray analysis. The complex crystallised in the orthorhombic space group P2(1)2(1)2(1) with cell dimensions a = 26.3 A, b = 52.0 A and c = 67.1 A, incorporating two B-DNA duplexes and four nogalamycin molecules in the asymmetric unit. The final refined structure included 97 water molecules, one spermine molecule, two acetate ions and one sodium ion, yielding an overall R factor of 19.2% (calculated using all 12,358 reflections in the resolution range 10.7 to 1.6 A) and an Rtree of 23.7% (using 1229 test reflections). The d5'(TGTACA)2 sequence was designed to include the d5'(TpG) pyrimidine-purine base step that has been ascertained as a preferential intercalation site. The complexes in the asymmetric unit are globally similar; one nogalamycin molecule intercalates between each d5'(TpG) step in each duplex. The DNA of each complex exists as a distorted B-DNA duplex displaying some Z-DNA character in the form of C3' endo sugars at some residues. Structural comparisons between the d5'(TGTACA)2-nogalamycin2 complex and the complexes of this drug with the sequences d5'(TGATCA)2 and d5'(5MeCGT(pS)A5MeCG)2 highlight differences in binding interactions between nogalamycin and these various triplet DNA binding sites, with regards to the stability of drug intercalation, which in turn is correlated to effective levels of cytotoxicity towards tumour cells. The number of both direct and water-mediated hydrogen bonds and van der Waal's interactions between substituents of nogalamycin and the d5'(TGTACA)2 and d5'(5MeCGT(pS)A5MeCG)2 sequences are significantly greater than those made with the d5'(TGATCA)2 sequence, suggesting that the central d5'(TpA) in the former confers additional stability to the complex once the drug has bound.
- Department of Chemistry, University of York, Heslington, England.
Organizational Affiliation: