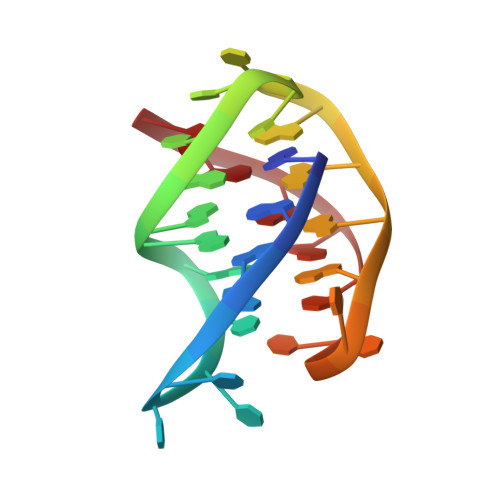
Solution structure of the Oxytricha telomeric repeat d[G4(T4G4)3] G-tetraplex.
Wang, Y., Patel, D.J.(1995) J Mol Biology 251: 76-94
- PubMed: 7643391 Search on PubMed
- DOI: https://doi.org/10.1006/jmbi.1995.0417
- Primary Citation Related Structures:
201D - PubMed Abstract:
The solution structure of Oxytricha telomere sequence d[G4(T4G4)3] in 0.1 M Na+ containing solution has been determined using a combined NMR-molecular dynamics approach including relaxation matrix refinement. This four G4 repeat sequence folds intramolecularly into a right-handed G-tetraplex containing four stacked G-tetrads which are connected by two lateral T4 loops and a central diagonal T4 loop. The guanine glycosidic bonds adopt a syn-anti alternation along the full length of the d[G4(T4G4)3] sequence while the orientation around adjacent G-tetrads switches between syn.syn.anti.anti and anti.anti.syn.syn alignments. Four distinct grooves are formed by the parallel (two of medium width) and anti-parallel (one wide and one narrow width) alignment of adjacent G-G-G-G segments in the G-tetraplex. The T4 residues in the diagonal loop are well-defined while the T4 residues in both lateral loops are under-defined and sample multiple conformations. The solution structure of the Na(+)-stabilized Oxytricha d[G4(T4G4)3] G-tetraplex and an earlier solution structure reported from our laboratory on the Na(+)-stabilized human d[AG3(T2AG3)3] G-tetraplex exhibit a common folding topology defined by the same syn/anti distribution of guanine residues along individual strands and around individual G-tetrads, as well as a common central diagonal loop which defines the strand directionalities. The well-resolved proton NMR spectra associated with the d[G4(T4G4)3] G-tetraplex opens the opportunity for studies ranging from cation-dependent characterization of G-tetraplex conformation and hydration to ligand and protein recognition of the distinct grooves associated with this folding topology.
- Cellular Biochemistry and Biophysics Program, Memorial Sloan-Kettering Cancer Center, New York, NY 10027, USA.
Organizational Affiliation: