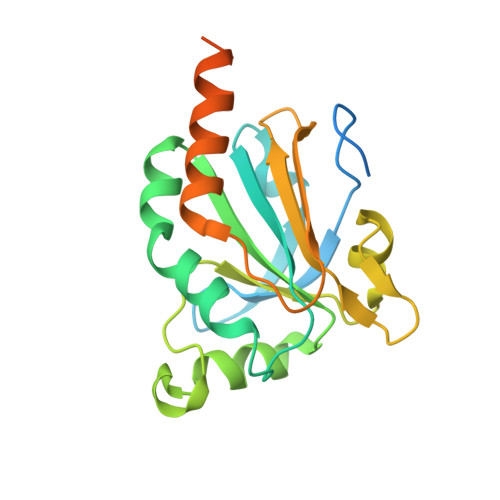
Bovine Mitochondrial Peroxiredoxin III Forms a Two-Ring Catenane
Cao, Z., Roszak, A.W., Gourlay, L.J., Lindsay, J.G., Isaacs, N.W.(2005) Structure 13: 1661-1664
- PubMed: 16271889
- DOI: https://doi.org/10.1016/j.str.2005.07.021
- Primary Citation Related Structures:
1ZYE - PubMed Abstract:
A crystal structure is reported for the C168S mutant of a typical 2-Cys peroxiredoxin III (Prx III) from bovine mitochondria at a resolution of 3.3 A. Prx III is present as a two-ring catenane comprising two interlocking dodecameric toroids that are assembled from basic dimeric units. Each ring has an external diameter of 150 A and encompasses a central cavity that is 70 A in width. The concatenated dodecamers are inclined at an angle of 55 degrees, which provides a large contact surface between the rings. Dimer-dimer contacts involved in toroid formation are hydrophobic in nature, whereas the 12 areas of contact between interlocked rings arise from polar interactions. These two major modes of subunit interaction provide important insights into possible mechanisms of catenane formation.
- Department of Biochemistry and Molecular Biology, University of Glasgow, Glasgow G12 8QQ, United Kingdom.
Organizational Affiliation: