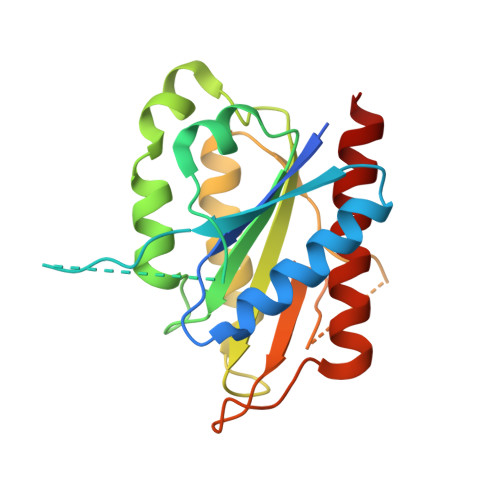
Crystal structures of the tryptophan repressor binding protein WrbA and complexes with flavin mononucleotide.
Gorman, J., Shapiro, L.(2005) Protein Sci 14: 3004-3012
- PubMed: 16322580
- DOI: https://doi.org/10.1110/ps.051680805
- Primary Citation Related Structures:
1YDG, 1YRH, 1ZWK, 1ZWL - PubMed Abstract:
The tryptophan repressor binding protein WrbA binds to the tryptophan repressor protein TrpR. Although the biological role of WrbA remains unclear, it has been proposed to function in enhancing the stability of TrpR-DNA complexes. Sequence database analysis has identified WrbA as a founding member of a flavodoxin-like family of proteins. Here we present crystal structures of WrbA from Deinococcus radiodurans and Pseudomonas aeruginosa and their complexes with flavin mononucleotide. The protomer structure is similar to that of previously determined long-chain flavodoxins; however, each contains a conserved inserted region unique to the WrbA family. Interestingly, each WrbA protein forms a homotetramer with 222 symmetry, unique among flavodoxin-like proteins, in which each protomer binds one flavin mononucleotide cofactor molecule.
- Department of Biochemistry and Molecular Biophysics, Columbia University, 630 West 168th Street, Box 18, New York, NY 10032, USA.
Organizational Affiliation: