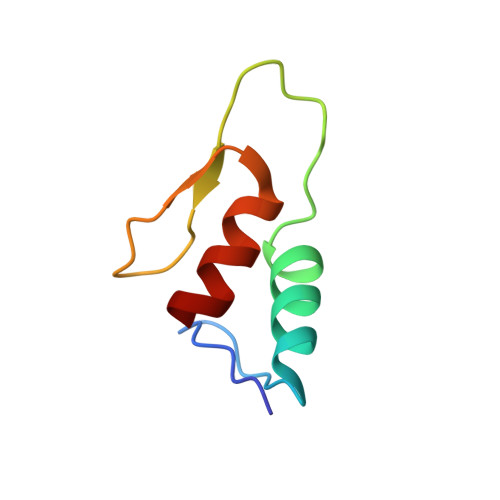
Solution structure of a Zap1 zinc-responsive domain provides insights into metalloregulatory transcriptional repression in Saccharomyces cerevisiae.
Wang, Z., Feng, L.S., Matskevich, V., Venkataraman, K., Parasuram, P., Laity, J.H.(2006) J Mol Biology 357: 1167-1183
- PubMed: 16483601
- DOI: https://doi.org/10.1016/j.jmb.2006.01.010
- Primary Citation of Related Structures:
1ZW8 - PubMed Abstract:
The Zap1 transcription factor controls expression of genes that regulate zinc homeostasis in Saccharomyces cerevisiae. The solution structure of two zinc fingers (zf1-2(CA3)) derived from a zinc-responsive domain of Zap1 (zf1-2) has been determined. Under zinc-limiting conditions, zinc finger 2 (zf2) from this domain has been shown to be a constitutive transcriptional activator. Moreover, repression of zf2 function in zinc-replete cells required zinc coordination to both canonical finger 1 (zf1) and zf2 metal sites, suggesting zf1-zf2 cooperativity underlies Zap1 metalloregulation. A structural basis for this cooperativity is identified here. Favorable inter-helical contacts in zf1-2(CA3) extend the individual finger hydrophobic cores through the zf1-zf2 interface. Tryptophan residues at position 5 in each finger provide numerous non-helical inter-finger contacts reminiscent of those observed in GLI1 zinc fingers 1 and 2. The molecular mechanism for zf1-dependent repression of zf2 transcriptional activation is explored further using NMR and CD titration studies. While zf1 independently forms a betabetaalpha solution structure, the majority of zf2 ensemble solution states do not adopt the canonical betabetaalpha zinc finger fold without zf1-zf2 interactions. Cooperative effects on Zn(II) affinities stemming from these finger-finger interactions are observed also in calorimetric studies, in which the 160(+/-20)nM (zf1) and 250(+/-40)nM (zf2) K(d) values for each individual finger increased substantially in the context of the zf1-2 protein (apparent K(dzf1-2WT)=4.6(+/-1.2)nM). On the basis of the above observations, we propose a mechanism for Zap1 transcriptional regulation in which zf1-zf2 interactions stabilize the betabetaalpha folded "repressed state" of the zf2 activation domain in the presence of cellular Zn(II) excess. Moreover, in contrast to earlier reports of <<1 labile zinc ion/Escherichia coli cell, the zf1-zf2 zinc affinities determined calorimetrically are consistent with Zn(II) levels >>1 labile zinc ion/eukaryotic cell.
- Division of Cell Biology and Biophysics, School of Biological Sciences, University of Missouri-Kansas City, Kansas City, MO 64110-2499, USA.
Organizational Affiliation: