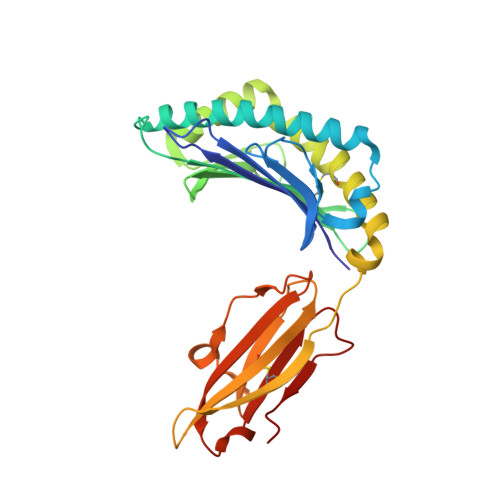
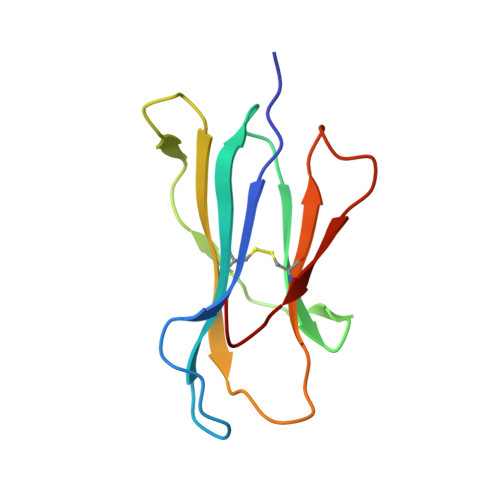
CTL Recognition of a Bulged Viral Peptide Involves Biased TCR Selection.
Miles, J.J., Elhassen, D., Borg, N.A., Silins, S.L., Tynan, F.E., Burrows, J.M., Purcell, A.W., Kjer-Nielsen, L., Rossjohn, J., Burrows, S.R., McCluskey, J., El-Hassen, D.(2005) J Immunol 175: 3826-3834
- PubMed: 16148129 Search on PubMed
- DOI: https://doi.org/10.4049/jimmunol.175.6.3826
- Primary Citation Related Structures:
1ZSD - PubMed Abstract:
MHC class I molecules generally present peptides of 8-10 aa long, forming an extended coil in the HLA cleft. Although longer peptides can also bind to class I molecules, they tend to bulge from the cleft and it is not known whether the TCR repertoire has sufficient plasticity to recognize these determinants during the antiviral CTL response. In this study, we show that unrelated individuals infected with EBV generate a significant CTL response directed toward an HLA-B*3501-restricted, 11-mer epitope from the BZLF1 Ag. The 11-mer determinant adopts a highly bulged conformation with seven of the peptide side chains being solvent-exposed and available for TCR interaction. Such a complex potentially creates a structural challenge for TCR corecognition of both HLA-B*3501 and the peptide Ag. Surprisingly, unrelated B*3501 donors recognizing the 11-mer use identical or closely related alphabeta TCR sequences that share particular CDR3 motifs. Within the small number of dominant CTL clonotypes observed, each has discrete fine specificity for the exposed side chain residues of the peptide. The data show that bulged viral peptides are indeed immunogenic but suggest that the highly constrained TCR repertoire reflects a limit to TCR diversity when responding to some unusual MHC peptide ligands.
- Cellular Immunology Laboratory, Queensland Institute of Medical Research, Brisbane, Australia.
Organizational Affiliation: