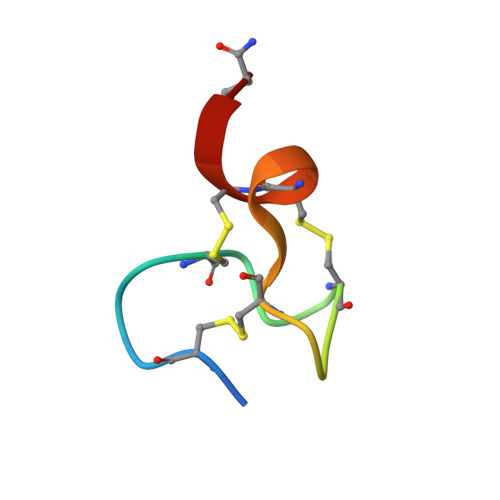
The Two Cysteine-rich Head Domains of Minicollagen from Hydra Nematocysts Differ in their Cystine Framework and Overall Fold Despite an Identical Cysteine Sequence Pattern.
Milbradt, A.G., Boulegue, C., Moroder, L., Renner, C.(2005) J Mol Biology 354: 591-600
- PubMed: 16257007
- DOI: https://doi.org/10.1016/j.jmb.2005.09.080
- Primary Citation of Related Structures:
1ZPX - PubMed Abstract:
Synthetic replicates of naturally occurring cysteine-rich peptides such as hormones, neurotransmitters, growth factors, enzyme inhibitors, defensins and toxins often can be oxidatively folded in high yields to their native structure in simple redox buffers. Thereby, identical cysteine patterns in the sequence were found to generate identical disulfide connectivities and homologous spatial structures despite significant variability in the non-cysteine positions. Minicollagen-1 from the nematocysts of Hydra is a trimeric protein that contains cysteine-rich domains at the N and C termini, which are involved in the assembly of an intermolecular disulfide network. Determination of the three-dimensional structures of peptides corresponding to the N-terminal and C-terminal domains by NMR spectroscopy revealed a remarkable exception from the general rule. Despite an identical cysteine pattern, the two domains of minicollagen-1 form different disulfide bridges and exhibit distinctly different folds, both of which are not found in the current structural databases. To our knowledge, this is the first case where two relatively short peptides with the abundant cysteine residues in identical sequence positions fold uniquely and with high yields into defined, but differing, structures. Therefore, the cysteine-rich domains of minicollagen constitute ideal model systems for studies of the interplay between folding and oxidation in proteins.
- Max-Planck-Institut für Biochemie, 82152 Martinsried, Germany.
Organizational Affiliation: