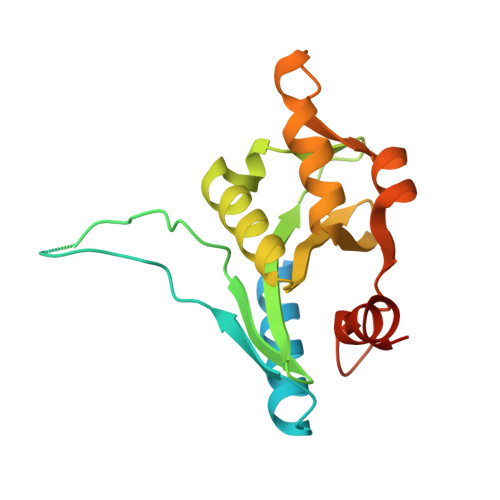
The Structure of Bacillus subtilis RecU Holliday Junction Resolvase and Its Role in Substrate Selection and Sequence-Specific Cleavage.
McGregor, N., Ayora, S., Sedelnikova, S., Carrasco, B., Alonso, J.C., Thaw, P., Rafferty, J.(2005) Structure 13: 1341-1351
- PubMed: 16154091 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.05.011
- Primary Citation Related Structures:
1ZP7 - PubMed Abstract:
We have determined the structure of the enzyme RecU from Bacillus subtilis, that is the general Holliday junction resolving enzyme in Gram-positive bacteria. The enzyme fold reveals a striking similarity to a class of resolvase enzymes found in archaeal sources and members of the type II restriction endonuclease family to which they are related. The structure confirms the presence of active sites formed around clusters of acidic residues that we have also shown to bind divalent cations. Mutagenesis data presented here support the key role of certain residues. The RecU structure suggests a basis for Holliday junction selectivity and suggests how sequence-specific cleavage might be achieved. Models for a resolvase-DNA complex address how the enzyme might organize junctions into an approximately 4-fold symmetric form.
- The Krebs Institute, Department of Molecular Biology and Biotechnology, University of Sheffield, Western Bank, Sheffield S10 2TN, United Kingdom.
Organizational Affiliation: