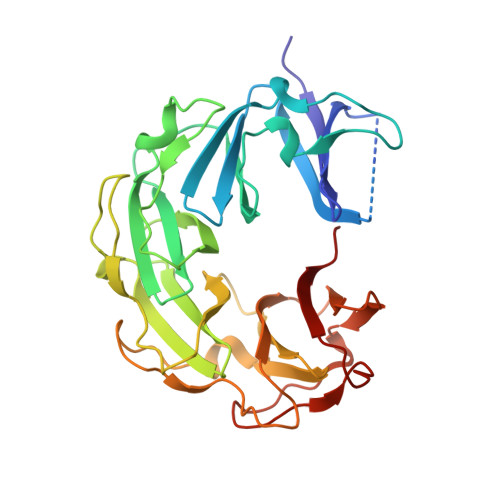
A superhelical spiral in the Escherichia coli DNA gyrase A C-terminal domain imparts unidirectional supercoiling bias
Ruthenburg, A.J., Graybosch, D.M., Huetsch, J.C., Verdine, G.L.(2005) J Biological Chem 280: 26177-26184
- PubMed: 15897198 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M502838200
- Primary Citation Related Structures:
1ZI0 - PubMed Abstract:
DNA gyrase is unique among type II topoisomerases in that its DNA supercoiling activity is unidirectional. The C-terminal domain of the gyrase A subunit (GyrA-CTD) is required for this supercoiling bias. We report here the x-ray structure of the Escherichia coli GyrA-CTD (Protein Data Bank code 1ZI0). The E. coli GyrA-CTD adopts a circular-shaped beta-pinwheel fold first seen in the Borrelia burgdorferi GyrA-CTD. However, whereas the B. burgdorferi GyrA-CTD is flat, the E. coli GyrA-CTD is spiral. DNA relaxation assays reveal that the E. coli GyrA-CTD wraps DNA inducing substantial (+) superhelicity, while the B. burgdorferi GyrA-CTD introduces a more modest (+) superhelicity. The observation of a superhelical spiral in the present structure and that of the Bacillus stearothermophilus ParC-CTD structure suggests unexpected similarities in substrate selectivity between gyrase and Topo IV enzymes. We propose a model wherein the right-handed ((+) solenoidal) wrapping of DNA around the E. coli GyrA-CTD enforces unidirectional (-) DNA supercoiling.
- Department of Chemistry and Chemical Biology, Harvard University, Cambridge, Massachusetts 02138, USA.
Organizational Affiliation: