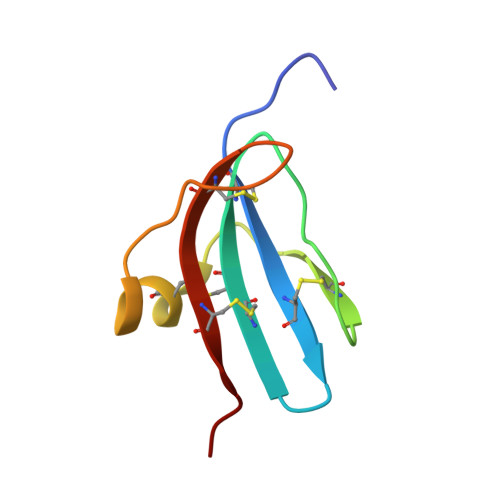
NMR structural characterization and computational predictions of the major intermediate in oxidative folding of leech carboxypeptidase inhibitor
Arolas, J.L., D'Silva, L., Popowicz, G.M., Aviles, F.X., Holak, T.A., Ventura, S.(2005) Structure 13: 1193-1202
- PubMed: 16084391 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2005.05.008
- Primary Citation Related Structures:
1ZFI, 1ZFL - PubMed Abstract:
The III-A intermediate constitutes the major rate-determining step in the oxidative folding of leech carboxypeptidase inhibitor (LCI). In this work, III-A has been directly purified from the folding reaction and structurally characterized by NMR spectroscopy. This species, containing three native disulfides, displays a highly native-like structure; however, it lacks some secondary structure elements, making it more flexible than native LCI. III-A represents a structurally determined example of a disulfide-insecure intermediate; direct oxidation of this species to the fully native protein seems to be restricted by the burial of its two free cysteine residues inside a native-like structure. We also show that theoretical approaches based on topological constraints predict with good accuracy the presence of this folding intermediate. Overall, the derived results suggest that, as it occurs with non-disulfide bonded proteins, native-like interactions between segments of secondary structure rather than the crosslinking of disulfide bonds direct the folding of LCI.
- Institut de Biotecnologia i de Biomedicina and Departament de Bioquímica i Biologia Molecular, Universitat Autònoma de Barcelona, 08193 Bellaterra, Spain.
Organizational Affiliation: