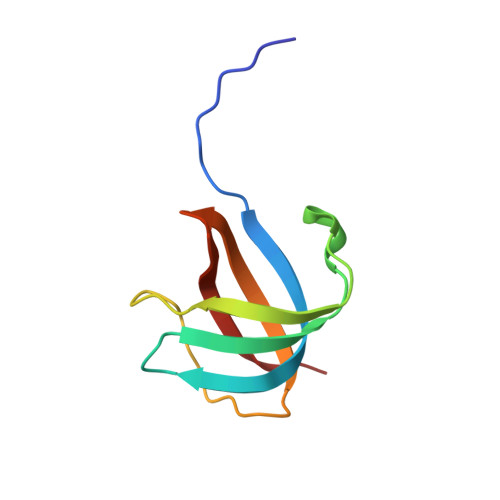
A Novel Copper-Binding Fold for the Periplasmic Copper Resistance Protein CusF.
Loftin, I.R., Franke, S., Roberts, S.A., Weichsel, A., Montfort, W.R., Rensing, C., McEvoy, M.M.(2005) Biochemistry 44: 10533-10540
- PubMed: 16060662 Search on PubMed
- DOI: https://doi.org/10.1021/bi050827b
- Primary Citation Related Structures:
1ZEQ - PubMed Abstract:
We have determined the crystal structure of apo-CusF, a periplasmic protein involved in copper and silver resistance in Escherichia coli. The protein forms a five-stranded beta-barrel, classified as an OB-fold, which is a unique topology for a copper-binding protein. NMR chemical shift mapping experiments suggest that Cu(I) is bound by conserved residues H36, M47, and M49 located in beta-strands 2 and 3. These residues are clustered at one end of the beta-barrel, and their side chains are oriented toward the interior of the barrel. Cu(I) can be modeled into the apo-CusF structure with only minimal structural changes using H36, M47, and M49 as ligands. The unique structure and metal binding site of CusF are distinct from those of previously characterized copper-binding proteins.
- Department of Biochemistry and Molecular Biophysics, University of Arizona, Tucson, Arizona 85721, USA.
Organizational Affiliation: