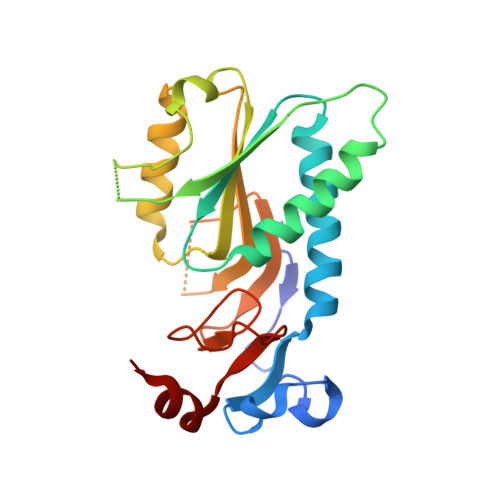
The Crystal Structure of Free Human Hypoxanthine-guanine Phosphoribosyltransferase Reveals Extensive Conformational Plasticity Throughout the Catalytic Cycle
Keough, D.T., Brereton, I.M., de Jersey, J., Guddat, L.W.(2005) J Mol Biology 351: 170-181
- PubMed: 15990111 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.05.061
- Primary Citation Related Structures:
1Z7G - PubMed Abstract:
Human hypoxanthine-guanine phosphoribosyltransferase (HGPRT) catalyses the synthesis of the purine nucleoside monophosphates, IMP and GMP, by the addition of a 6-oxopurine base, either hypoxanthine or guanine, to the 1-beta-position of 5-phospho-alpha-d-ribosyl-1-pyrophosphate (PRib-PP). The mechanism is sequential, with PRib-PP binding to the free enzyme prior to the base. After the covalent reaction, pyrophosphate is released followed by the nucleoside monophosphate. A number of snapshots of the structure of this enzyme along the reaction pathway have been captured. These include the structure in the presence of the inactive purine base analogue, 7-hydroxy [4,3-d] pyrazolo pyrimidine (HPP) and PRib-PP.Mg2+, and in complex with IMP or GMP. The third structure is that of the immucillinHP.Mg(2+).PP(i) complex, a transition-state analogue. Here, the first crystal structure of free human HGPRT is reported to 1.9A resolution, showing that significant conformational changes have to occur for the substrate(s) to bind and for catalysis to proceed. Included in these changes are relative movement of subunits within the tetramer, rotation and extension of an active-site alpha-helix (D137-D153), reorientation of key active-site residues K68, D137 and K165, and the rearrangement of three active-site loops (100-128, 165-173 and 186-196). Toxoplasma gondii HGXPRT is the only other 6-oxopurine phosphoribosyltransferase structure solved in the absence of ligands. Comparison of this structure with human HGPRT reveals significant differences in the two active sites, including the structure of the flexible loop containing K68 (human) or K79 (T.gondii).
- School of Molecular and Microbial Sciences, University of Queensland, Brisbane 4072, Australia.
Organizational Affiliation: