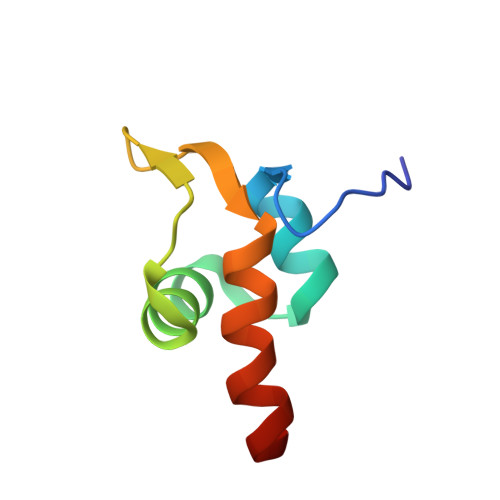
Structural and genetic analyses reveal a key role in prophage excision for the TorI response regulator inhibitor
Elantak, L., Ansaldi, M., Guerlesquin, F., Mejean, V., Morelli, X.(2005) J Biological Chem 280: 36802-36808
- PubMed: 16079126
- DOI: https://doi.org/10.1074/jbc.M507409200
- Primary Citation of Related Structures:
1Z4H - PubMed Abstract:
TorI (Tor inhibition protein) has been identified in Escherichia coli as a protein inhibitor acting through protein-protein interaction with the TorR response regulator. This interaction, which does not interfere with TorR DNA binding activity, probably prevents the recruitment of RNA polymerase to the torC promoter. In this study we have solved the solution structure of TorI, which adopts a prokaryotic winged-helix arrangement. Despite no primary sequence similarity, the three-dimensional structure of TorI is highly homologous to the (lambda)Xis, Mu bacteriophage repressor (MuR-DBD), and transposase (MuA-DBD) structures. We propose that the TorI protein is the structural missing link between the (lambda)Xis and MuR proteins. Moreover, in vivo assays demonstrated that TorI plays an essential role in prophage excision. Heteronuclear NMR experiments and site-directed mutagenesis studies have pinpointed out key residues involved in the DNA binding activity of TorI. Our findings suggest that TorI-related proteins identified in various pathogenic bacterial genomes define a new family of atypical excisionases.
- Unité de Bioénergétique et Ingénierie des Protéines, IBSM-CNRS, 31 chemin Joseph Aiguier, 13402 Marseille Cedex 20, France.
Organizational Affiliation: