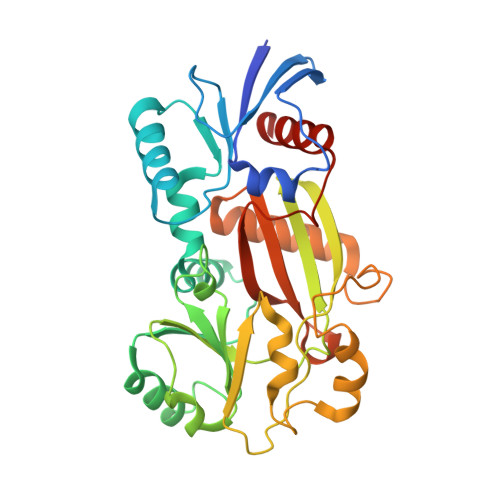
Specificity determinants in inositol polyphosphate synthesis: crystal structure of inositol 1,3,4-trisphosphate 5/6-kinase.
Miller, G.J., Wilson, M.P., Majerus, P.W., Hurley, J.H.(2005) Mol Cell 18: 201-212
- PubMed: 15837423 Search on PubMed
- DOI: https://doi.org/10.1016/j.molcel.2005.03.016
- Primary Citation Related Structures:
1Z2N, 1Z2O, 1Z2P - PubMed Abstract:
Inositol hexakisphosphate and other inositol high polyphosphates have diverse and critical roles in eukaryotic regulatory pathways. Inositol 1,3,4-trisphosphate 5/6-kinase catalyzes the rate-limiting step in inositol high polyphosphate synthesis in animals. This multifunctional enzyme also has inositol 3,4,5,6-tetrakisphosphate 1-kinase and other activities. The structure of an archetypal family member, from Entamoeba histolytica, has been determined to 1.2 A resolution in binary and ternary complexes with nucleotide, substrate, and product. The structure reveals an ATP-grasp fold. The inositol ring faces ATP edge-on such that the 5- and 6-hydroxyl groups are nearly equidistant from the ATP gamma-phosphate in catalytically productive phosphoacceptor positions and explains the unusual dual site specificity of this kinase. Inositol tris- and tetrakisphosphates interact via three phosphate binding subsites and one solvent-exposed site that could in principle be occupied by 18 different substrates, explaining the mechanisms for the multiple specificities and catalytic activities of this enzyme.
- Laboratory of Molecular Biology, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, United States Department of Health and Human Services, Bethesda, Maryland 20892, USA.
Organizational Affiliation: