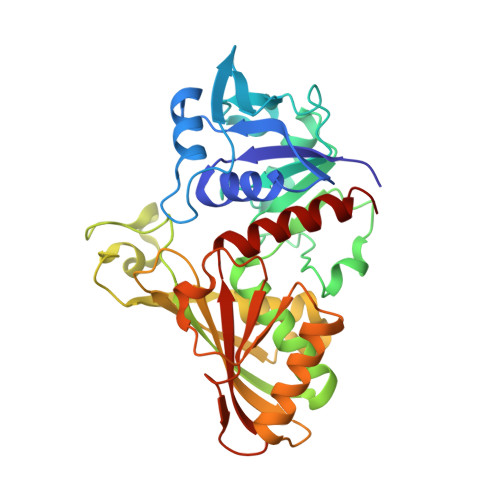
Structure of glyceraldehyde-3-phosphate dehydrogenase from Plasmodium falciparum.
Satchell, J.F., Malby, R.L., Luo, C.S., Adisa, A., Alpyurek, A.E., Klonis, N., Smith, B.J., Tilley, L., Colman, P.M.(2005) Acta Crystallogr D Biol Crystallogr 61: 1213-1221
- PubMed: 16131754
- DOI: https://doi.org/10.1107/S0907444905018317
- Primary Citation of Related Structures:
1YWG - PubMed Abstract:
The malaria parasite Plasmodium falciparum is responsible for about two million deaths annually, making it important to obtain information about enzymes from this organism that represent potential drug targets. The gene for P. falciparum glyceraldehyde-3-phosphate dehydrogenase (PfGAPDH) has been cloned and the protein expressed as a hexahistidine-tagged recombinant protein in Escherichia coli. The recombinant protein has been crystallized and its three-dimensional structure determined. One molecule of the cofactor NAD+ is bound to each of the four subunits in the tetrameric enzyme. The major structural feature distinguishing human GAPDH from PfGAPDH is the insertion of a dipeptide (-KG-) in the so-called S loop. This insert, together with other characteristic single-amino-acid substitutions, alters the chemical environment of the groove that encompasses the R dyad and that links adjacent cofactor-binding sites and may be responsible for the selective inhibition of the enzyme by ferriprotoporphyrin IX.
- The Walter and Eliza Hall Institute of Medical Research, Melbourne, Victoria, Australia.
Organizational Affiliation: