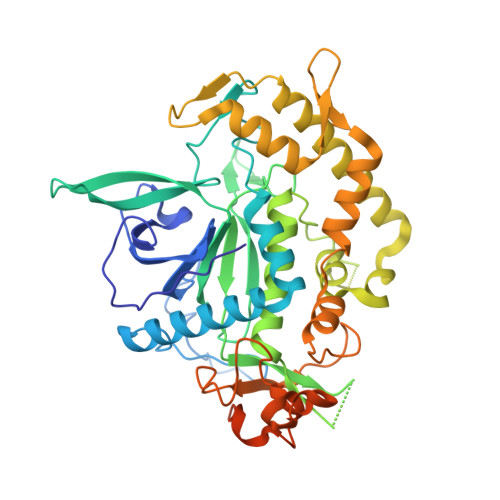
Structural analysis of the catalytic domain of tetanus neurotoxin.
Rao, K.N., Kumaran, D., Binz, T., Swaminathan, S.(2005) Toxicon 45: 929-939
- PubMed: 15904688 Search on PubMed
- DOI: https://doi.org/10.1016/j.toxicon.2005.02.032
- Primary Citation Related Structures:
1YVG - PubMed Abstract:
Clostridium neurotoxins, comprising the tetanus neurotoxin and the seven antigenically distinct botulinum neurotoxins (BoNT/A-G), are among the known most potent bacterial protein toxins to humans. Although they have similar function, sequences and three-dimensional structures, the substrate specificity and the selectivity of peptide bond cleavage are different and unique. Tetanus and botulinum type B neurotoxins enzymatically cleave the same substrate, vesicle-associated membrane protein, at the same peptide bond though the optimum length of substrate peptide required for cleavage by them is different. Here, we present the first experimentally determined three-dimensional structure of the catalytic domain of tetanus neurotoxin and analyze its active site. The structure provides insight into the active site of tetanus toxin's proteolytic activity and the importance of the nucleophilic water and the role of the zinc ion. The probable reason for different modes of binding of vesicle-associated membrane protein to botulinum neurotoxin type B and the tetanus toxin is discussed. The structure provides a basis for designing a novel recombinant vaccine or structure-based drugs for tetanus.
- Brookhaven National Laboratory, Biology Department, Upton, NY 11973, USA.
Organizational Affiliation: