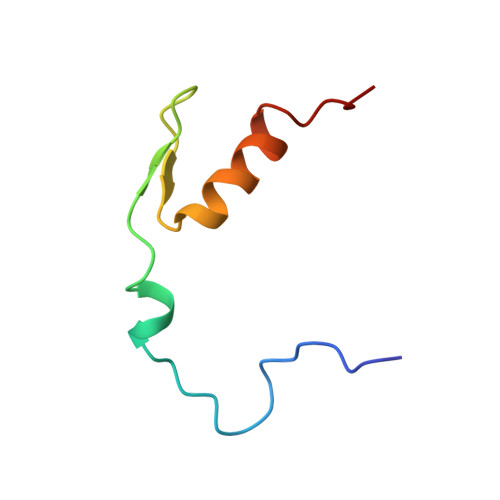
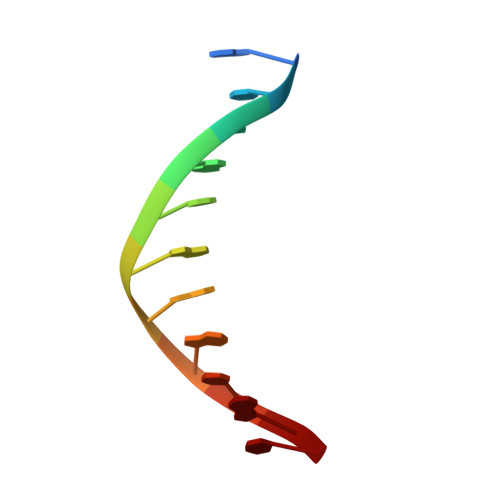
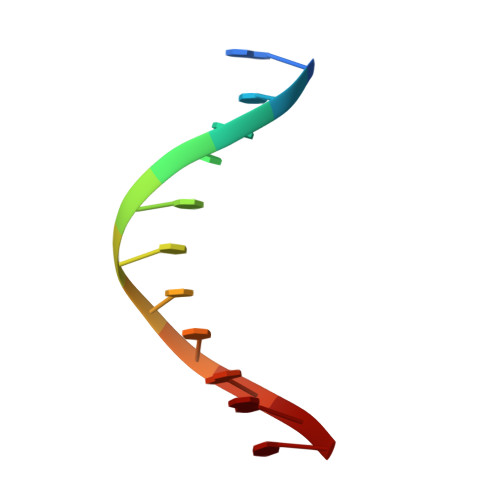
The solution structure of a specific GAGA factor-DNA complex reveals a modular binding mode.
Omichinski, J.G., Pedone, P.V., Felsenfeld, G., Gronenborn, A.M., Clore, G.M.(1997) Nat Struct Biol 4: 122-132
- PubMed: 9033593 Search on PubMed
- DOI: https://doi.org/10.1038/nsb0297-122
- Primary Citation Related Structures:
1YUI, 1YUJ - PubMed Abstract:
The structure of a complex between the DNA binding domain of the GAGA factor (GAGA-DBD) and an oligonucleotide containing its GAGAG consensus binding site has been determined by nuclear magnetic resonance spectroscopy. The GAGA-DBD comprises a single classical Cys2-His2 zinc finger core, and an N-terminal extension containing two highly basic regions, BR1 and BR2. The zinc finger core binds in the major groove and recognizes the first three GAG bases of the consensus in a manner similar to that seen in other classical zinc finger-DNA complexes. Unlike the latter, which require tandem zinc finger repeats with a minimum of two units for high affinity binding, the GAGA-DBD makes use of only a single finger complemented by BR1 and BR2. BR2 forms a helix that interacts in the major groove recognizing the last G of the consensus, while BR1 wraps around the DNA in the minor groove and recognizes the A in the fourth position of the consensus. The implications of the structure of the GAGA-DBD-DNA complex for chromatin remodelling are discussed.
- Laboratories of Chemical Physics, National Institute of Diabetes and Digestive and Kidney Diseases, National Institutes of Health, Bethesda, Maryland 20892-0520, USA.
Organizational Affiliation: