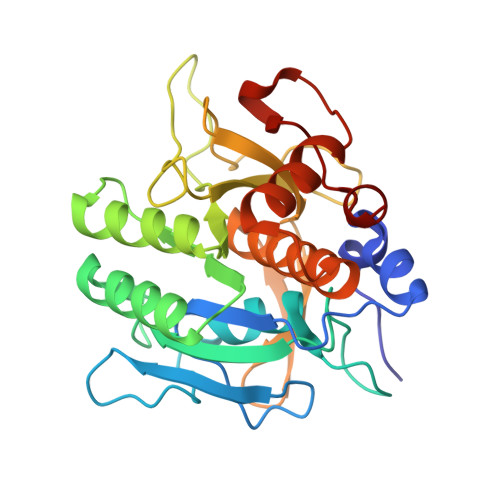
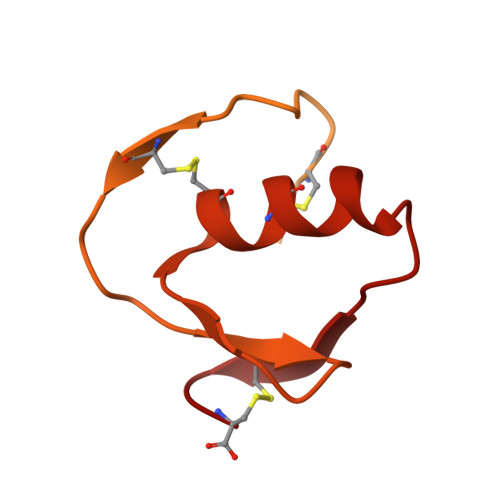
Structure of the subtilisin Carlsberg-OMTKY3 complex reveals two different ovomucoid conformations.
Maynes, J.T., Cherney, M.M., Qasim, M.A., Laskowski Jr, M., James, M.N.(2005) Acta Crystallogr D Biol Crystallogr 61: 580-588
- PubMed: 15858268 Search on PubMed
- DOI: https://doi.org/10.1107/S0907444905004889
- Primary Citation Related Structures:
1YU6 - PubMed Abstract:
One of the most studied protein proteinase inhibitors is the turkey ovomucoid third domain, OMTKY3. This inhibitor contains a reactive-site loop (Lys13I-Arg21I) that binds in a nearly identical manner to all studied serine proteinases, regardless of their clan or specificity. The crystal structure of OMTKY3 bound to subtilisin Carlsberg (CARL) has been determined. There are two complete copies of the complexes in the crystallographic asymmetric unit. Whereas the two enzyme molecules are virtually identical [0.16 A root-mean-square difference (r.m.s.d.) for 274 C(alpha) atoms], the two inhibitor molecules show dramatic differences between one another (r.m.s.d. = 2.4 A for 50 C(alpha) atoms). When compared with other proteinase-bound OMTKY3 molecules, these inhibitors show even larger differences. This work facilitates a re-evaluation of the importance of certain ovomucoid residues in proteinase binding and explains why additivity and sequence-based binding-prediction methods fail for the CARL-OMTKY3 complex.
- Canadian Institutes of Health Research, Group in Protein Structure and Function, Department of Biochemistry, Faculty of Medicine, University of Alberta, Edmonton, Alberta T6G 2H7, Canada. jason@biochem.ualberta.ca
Organizational Affiliation: