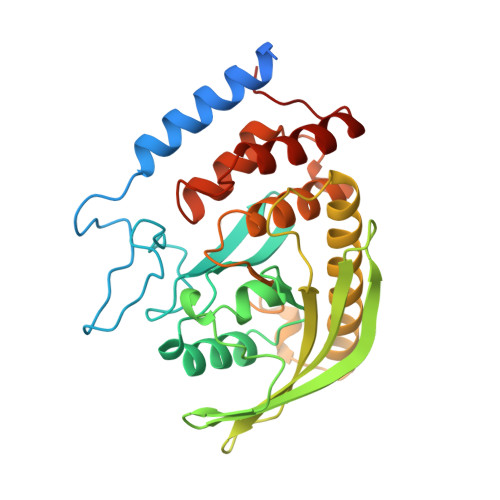
Crystal structure of Yersinia protein tyrosine phosphatase at 2.5 A and the complex with tungstate.
Stuckey, J.A., Schubert, H.L., Fauman, E.B., Zhang, Z.Y., Dixon, J.E., Saper, M.A.(1994) Nature 370: 571-575
- PubMed: 8052312 Search on PubMed
- DOI: https://doi.org/10.1038/370571a0
- Primary Citation Related Structures:
1YPT - PubMed Abstract:
Protein tyrosine phosphatases (PTPases) and kinases coregulate the critical levels of phosphorylation necessary for intracellular signalling, cell growth and differentiation. Yersinia, the causative bacteria of the bubonic plague and other enteric diseases, secrete an active PTPase, Yop51, that enters and suppresses host immune cells. Though the catalytic domain is only approximately 20% identical to human PTP1B, the Yersinia PTPase contains all of the invariant residues present in eukaryotic PTPases, including the nucleophilic Cys 403 which forms a phosphocysteine intermediate during catalysis. We present here structures of the unliganded (2.5 A resolution) and tungstate-bound (2.6 A) crystal forms which reveal that Cys 403 is positioned at the centre of a distinctive phosphate-binding loop. This loop is at the hub of several hydrogen-bond arrays that not only stabilize a bound oxyanion, but may activate Cys 403 as a reactive thiolate. Binding of tungstate triggers a conformational change that traps the oxyanion and swings Asp 356, an important catalytic residue, by approximately 6 A into the active site. The same anion-binding loop in PTPases is also found in the enzyme rhodanese.
- Department of Biological Chemistry, University of Michigan, Ann Arbor 48109-1055.
Organizational Affiliation: