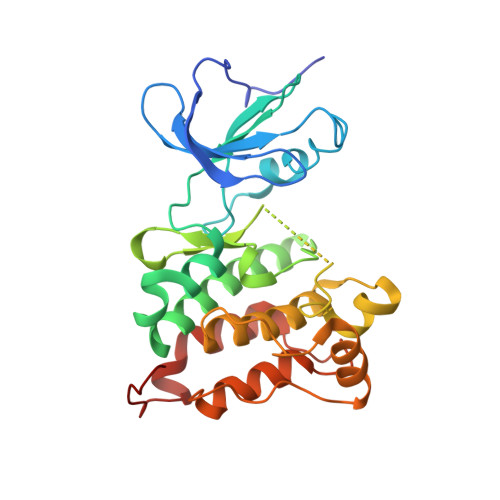
Crystal structures of active SRC kinase domain complexes
Breitenlechner, C.B., Kairies, N.A., Honold, K., Scheiblich, S., Koll, H., Greiter, E., Koch, S., Schaefer, W., Huber, R., Engh, R.A.(2005) J Mol Biology 353: 222-231
- PubMed: 16168436 Search on PubMed
- DOI: https://doi.org/10.1016/j.jmb.2005.08.023
- Primary Citation Related Structures:
1YOJ, 1YOL, 1YOM - PubMed Abstract:
c-Src was the first proto-oncoprotein to be identified, and has become the focus of many drug discovery programs. Src structures of a major inactive form have shown how the protein kinase is rigidified by several interdomain interactions; active configurations of Src are generated by release from this "assembled" or "bundled" form. Despite the importance of Src as a drug target, there is relatively little structural information available regarding the presumably more flexible active forms. Here we report three crystal structures of a dimeric active c-Src kinase domain, in an apo and two ligand complexed forms, with resolutions ranging from 2.9A to 1.95A. The structures show how the kinase domain, in the absence of the rigidifying interdomain interactions of the inactivation state, adopts a more open and flexible conformation. The ATP site inhibitor CGP77675 binds to the protein kinase with canonical hinge hydrogen bonds and also to the c-Src specific threonine 340. In contrast to purvalanol B binding in CDK2, purvalanol A binds in c-Src with a conformational change in a more open ATP pocket.
- Abteilung Strukturforschung, Max-Planck-Institut für Biochemie, 82152 Martinsried, Germany.
Organizational Affiliation: