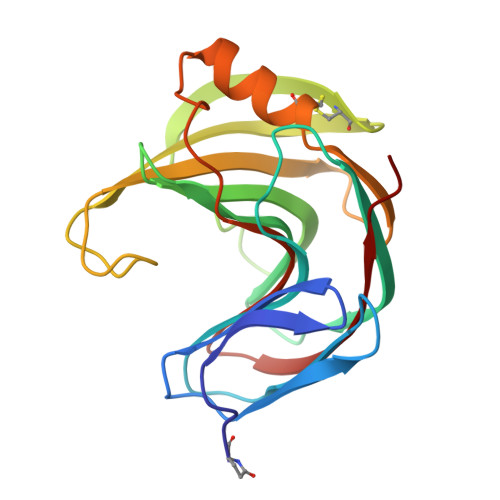
Thermophilic xylanase from Thermomyces lanuginosus: high-resolution X-ray structure and modeling studies.
Gruber, K., Klintschar, G., Hayn, M., Schlacher, A., Steiner, W., Kratky, C.(1998) Biochemistry 37: 13475-13485
- PubMed: 9753433 Search on PubMed
- DOI: https://doi.org/10.1021/bi980864l
- Primary Citation Related Structures:
1YNA - PubMed Abstract:
The crystal structure of the thermostable xylanase from Thermomyces lanuginosus was determined by single-crystal X-ray diffraction. The protein crystallizes in space group P21, a = 40.96(4) A, b = 52. 57(5) A, c = 50.47 (5) A, beta = 100.43(5) degrees, Z = 2. Diffraction data were collected at room temperature for a resolution range of 25-1.55 A, and the structure was solved by molecular replacement with the coordinates of xylanase II from Trichoderma reesei as a search model and refined to a crystallographic R-factor of 0.155 for all observed reflections. The enzyme belongs to the family 11 of glycosyl hydrolases [Henrissat, B., and Bairoch, A. (1993) Biochem. J. 293, 781-788]. pKa calculations were performed to assess the protonation state of residues relevant for catalysis and enzyme stability, and a heptaxylan was fitted into the active-site groove by homology modeling, using the published crystal structure of a complex between the Bacillus circulans xylanase and a xylotetraose. Molecular dynamics indicated the central three sugar rings to be tightly bound, whereas the peripheral ones can assume different orientations and conformations, suggesting that the enzyme might also accept xylan chains which are branched at these positions. The reasons for the thermostability of the T. lanuginosus xylanase were analyzed by comparing its crystal structure with known structures of mesophilic family 11 xylanases. It appears that the thermostability is due to the presence of an extra disulfide bridge, as well as to an increase in the density of charged residues throughout the protein.
- Institut für Physikalische Chemie, Universität Graz, Austria.
Organizational Affiliation: