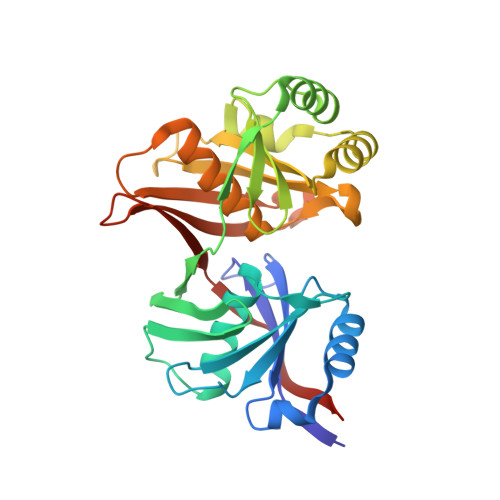
Crystal structure of YHI9, the yeast member of the phenazine biosynthesis PhzF enzyme superfamily
Liger, D., Quevillon-Cheruel, S., Sorel, I., Bremang, M., Blondeau, K., Aboulfath, I., Janin, J., van Tilbeurgh, H., Leulliot, N.(2005) Proteins 60: 778-786
- PubMed: 16021630 Search on PubMed
- DOI: https://doi.org/10.1002/prot.20548
- Primary Citation Related Structures:
1YM5 - PubMed Abstract:
In the Pseudomonas bacterial genomes, the PhzF proteins are involved in the production of phenazine derivative antibiotic and antifungal compounds. The PhzF superfamily however also encompasses proteins in all genomes from bacteria to eukaryotes, for which no function has been assigned. We have determined the three dimensional crystal structure at 2.05 A resolution of YHI9, the yeast member of the PhzF family. YHI9 has a fold similar to bacterial diaminopimelate epimerase, revealing a bimodular structure with an internal symmetry. Residue conservation identifies a putative active site at the interface between the two domains. Evolution of this protein by gene duplication, gene fusion and domain swapping from an ancestral gene containing the "hot dog" fold, identifies the protein as a "kinked double hot dog" fold.
- Institut de Biochimie et de Biophysique Moléculaire et Cellulaire (CNRS-UMR 8619), Université Paris-Sud, Orsay, France.
Organizational Affiliation: