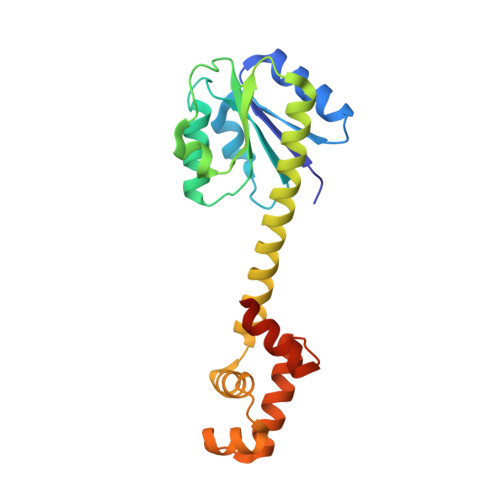
An Active-like Structure in the Unphosphorylated StyR Response Regulator Suggests a Phosphorylation- Dependent Allosteric Activation Mechanism
Milani, M., Leoni, L., Rampioni, G., Zennaro, E., Ascenzi, P., Bolognesi, M.(2005) Structure 13: 1289-1297
- PubMed: 16154086
- DOI: https://doi.org/10.1016/j.str.2005.05.014
- Primary Citation Related Structures:
1YIO, 1ZN2 - PubMed Abstract:
StyR belongs to the FixJ subfamily of signal transduction response regulators; it controls transcription of the styABCD operon coding for styrene catabolism in Pseudomonas fluorescens ST. The crystal structure of unphosphorylated StyR is reported at 2.2 A resolution. StyR is composed of an N-terminal regulatory domain (StyR-N) and a C-terminal DNA binding domain (StyR-C). The two domains are separated by an elongated linker alpha helix (34 residues), a new feature in known response regulator structures. StyR-C is structured similarly to the DNA binding domain of the response regulator NarL. StyR-N shows structural reorganization of the phosphate receiving region involved in activation/homodimerization: specific residues adopt an "active-like" conformation, and the alpha4 helix, involved in dimerization of the homologous FixJ response regulator, is trimmed to just one helical turn. Overall, structural considerations suggest that phosphorylation may act as an allosteric switch, shifting a preexisting StyR equilibrium toward the active, dimeric, DNA binding form.
- Giannina Gaslini Institute and INFM, Largo G. Gaslini 5, I-16147 Genova, Italy.
Organizational Affiliation: