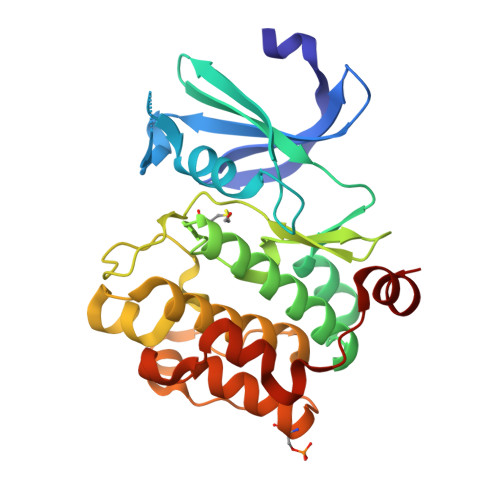
Pim-1 ligand-bound structures reveal the mechanism of serine/threonine kinase inhibition by LY294002.
Jacobs, M.D., Black, J., Futer, O., Swenson, L., Hare, B., Fleming, M., Saxena, K.(2005) J Biological Chem 280: 13728-13734
- PubMed: 15657054 Search on PubMed
- DOI: https://doi.org/10.1074/jbc.M413155200
- Primary Citation Related Structures:
1YHS, 1YI3, 1YI4 - PubMed Abstract:
Pim-1 is an oncogene-encoded serine/threonine kinase primarily expressed in hematopoietic and germ cell lines. Pim-1 kinase was originally identified in Maloney murine leukemia virus-induced T-cell lymphomas and is associated with multiple cellular functions such as proliferation, survival, differentiation, apoptosis, and tumorigenesis (Wang, Z., Bhattacharya, N., Weaver, M., Petersen, K., Meyer, M., Gapter, L., and Magnuson, N. S. (2001) J. Vet. Sci. 2, 167-179). The crystal structures of Pim-1 complexed with staurosporine and adenosine were determined. Although a typical two-domain serine/threonine protein kinase fold is observed, the inter-domain hinge region is unusual in both sequence and conformation; a two-residue insertion causes the hinge to bulge away from the ATP-binding pocket, and a proline residue in the hinge removes a conserved main chain hydrogen bond donor. Without this hydrogen bond, van der Waals interactions with the hinge serve to position the ligand. The hinge region of Pim-1 resembles that of phosphatidylinositol 3-kinase more closely than it does other protein kinases. Although the phosphatidylinositol 3-kinase inhibitor LY294002 also inhibits Pim-1, the structure of the LY294002.Pim-1 complex reveals a new binding mode that may be general for Ser/Thr kinases.
- Vertex Pharmaceuticals Incorporated, Cambridge, Massachusetts 02139, USA. marc_jacobs@vrtx.com
Organizational Affiliation: