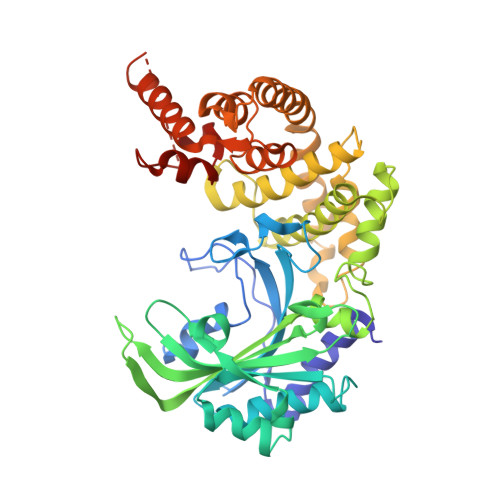
Breaking sieve for steric exclusion of a noncognate amino acid from active site of a tRNA synthetase.
Swairjo, M.A., Schimmel, P.R.(2005) Proc Natl Acad Sci U S A 102: 988-993
- PubMed: 15657145
- DOI: https://doi.org/10.1073/pnas.0409024102
- Primary Citation of Related Structures:
1YFR, 1YFS, 1YFT, 1YGB - PubMed Abstract:
The genetic code is fixed in aminoacylation reactions catalyzed by aminoacyl-tRNA synthetases. Amino acid discrimination occurs at two sites: one for amino acid activation and aminoacylation and one for editing misactivated amino acids. Although the active site sieves out bulkier amino acids, misactivation occurs with substrates whose side chains are smaller than the cognate one. Paradoxically, although alanyl-tRNA synthetase activates glycine as well as alanine, the sterically larger (than alanine) serine is also misactivated. Here, we report crystal structures of an active fragment of Aquifex aeolicus alanyl-tRNA synthetase complexed, separately, with Mg2+-ATP, alanine, glycine, and serine. Ala and Gly are bound in similar orientations in a side-chain-accommodating pocket, where alpha-amino and carboxyl groups are stabilized by salt bridges, and the carboxyl by an H-bond from the side chain NH2 of Asn-194. In contrast, whereas the same two salt bridges stabilize bound Ser, H-bonding of the highly conserved (among class II tRNA synthetases) Asn-194 side chain NH2 to the Ser OH, instead of to the carboxyl, forces pocket expansion. Significantly, in the Mg2+-ATP complex, Asn-194 coordinates a Mg2+-alpha-phosphate bridge. Thus, the sieve for Ser exclusion is broken because of selective pressure to retain Asn-194 for Mg2+-ATP and Ala binding.
- The Skaggs Institute for Chemical Biology and Department of Molecular Biology, The Scripps Research Institute, BCC-379, 10550 North Torrey Pines Road, La Jolla, CA 92037, USA.
Organizational Affiliation: