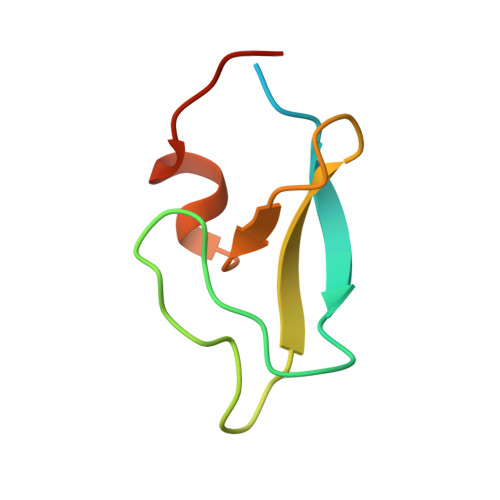
Intramolecular occlusion of the diacylglycerol-binding site in the C1 domain of munc13-1.
Shen, N., Guryev, O., Rizo, J.(2005) Biochemistry 44: 1089-1096
- PubMed: 15667202
- DOI: https://doi.org/10.1021/bi0476127
- Primary Citation of Related Structures:
1Y8F - PubMed Abstract:
Protein kinase C (PKC) isozymes and other receptors of diacylglycerol (DAG) bind to this widespread second messenger through their C(1) domains. These alternative DAG receptors include munc13-1, a large neuronal protein that is crucial for DAG-dependent augmentation of neurotransmitter release. Whereas the structures of several PKC C(1) domains have been determined and have been shown to require little conformational changes for ligand binding, it is unclear whether the C(1) domains from other DAG receptors contain specific structural features with key functional significance. To gain insight into this question, we have determined the three-dimensional structure in solution of the munc13-1 C(1) domain using NMR spectroscopy. The overall structure includes two beta-sheets, a short C-terminal alpha-helix, and two Zn(2+)-binding sites, resembling the structures of PKC C(1) domains. However, the munc13-1 C(1) domain exhibits striking structural differences with the PKC C(1) domains in the ligand-binding site. These differences result in occlusion of the binding site of the munc13-1 C(1) domain by a conserved tryptophan side chain that in PKCs adopts a completely different orientation. As a consequence, the munc13-1 C(1) domain requires a considerable conformational change for ligand binding. This structural distinction is expected to decrease the DAG affinity of munc13-1 compared to that of PKCs, and is likely to be critical for munc13-1 function. On the basis of these results, we propose that augmentation of neurotransmitter release may be activated at higher DAG levels than PKCs as a potential mechanism for uncoupling augmentation of release from the multitude of other signaling processes mediated by DAG.
- Department of Biochemistry, University of Texas Southwestern Medical Center, 5323 Harry Hines Boulevard, Dallas, Texas 75390, USA.
Organizational Affiliation: