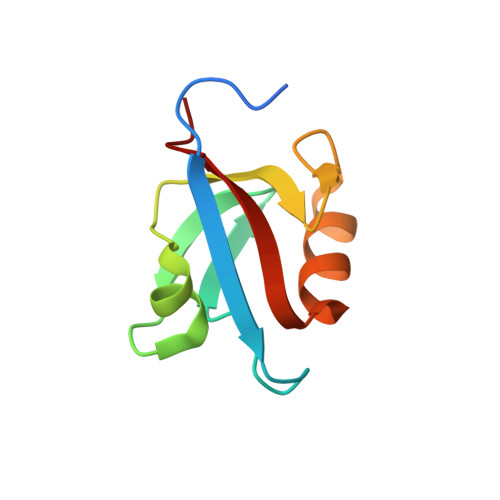
Solution structure of the second PDZ domain of the neuronal adaptor X11alpha and its interaction with the C-terminal peptide of the human copper chaperone for superoxide dismutase
Duquesne, A.E., de Ruijter, M., Brouwer, J., Drijfhout, J.W., Nabuurs, S.B., Spronk, C.A.E.M., Vuister, G.W., Ubbink, M., Canters, G.W.(2005) J Biomol NMR 32: 209-218
- PubMed: 16132821 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-005-7333-1
- Primary Citation Related Structures:
1Y7N - PubMed Abstract:
Protection against reactive oxygen species is provided by the copper containing enzyme superoxide dismutase 1 (SOD1). The copper chaperone CCS is responsible for copper insertion into apo-SOD1. This role is impaired by an interaction between the second PDZ domain (PDZ2alpha) of the neuronal adaptor protein X11alpha and the third domain of CCS (McLoughlin et al. (2001) J. Biol. Chem., 276, 9303-9307). The solution structure of the PDZ2alpha domain has been determined and the interaction with peptides derived from CCS has been explored. PDZ2alpha binds to the last four amino acids of the CCS protein (PAHL) with a dissociation constant of 91 +/- 2 microM. Peptide variants have been used to map the interaction areas on PDZ2alpha for each amino acid, showing an important role for the C-terminal leucine, in line with canonical PDZ-peptide interactions.
- Leiden Institute of Chemistry, Leiden University, The Netherlands.
Organizational Affiliation: