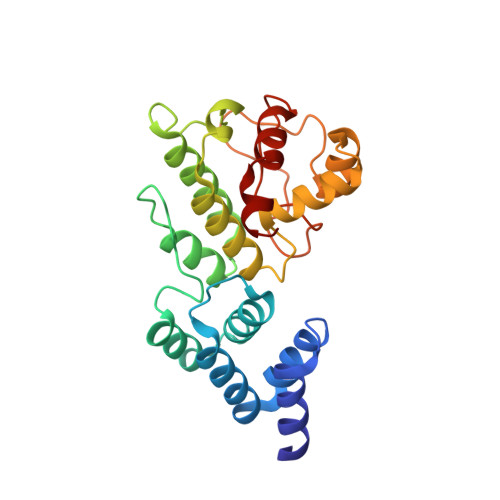
Structure of the Mg-chelatase cofactor GUN4 reveals a novel hand-shaped fold for porphyrin binding
Verdecia, M.A., Larkin, R.M., Ferrer, J.L., Riek, R., Chory, J., Noel, J.P.(2005) PLoS Biol 3: 151-151
- PubMed: 15884974 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1371/journal.pbio.0030151
- Primary Citation Related Structures:
1Y6I - PubMed Abstract:
In plants, the accumulation of the chlorophyll precursor Mg-protoporphyrin IX (Mg-Proto) in the plastid regulates the expression of a number of nuclear genes with functions related to photosynthesis. Analysis of the plastid-to-nucleus signaling activity of Mg-Proto in Arabidopsis thaliana led to the discovery of GUN4, a novel porphyrin-binding protein that also dramatically enhances the activity of Mg-chelatase, the enzyme that synthesizes Mg-Proto. GUN4 may also play a role in both photoprotection and the cellular shuttling of tetrapyrroles. Here we report a 1.78-A resolution crystal structure of Synechocystis GUN4, in which the porphyrin-binding domain adopts a unique three dimensional fold with a "cupped hand" shape. Biophysical and biochemical analyses revealed the specific site of interaction between GUN4 and Mg-Proto and the energetic determinants for the GUN4.Mg-Proto interaction. Our data support a novel protective function for GUN4 in tetrapyrrole trafficking. The combined structural and energetic analyses presented herein form the physical-chemical basis for understanding GUN4 biological activity, including its role in the stimulation of Mg-chelatase activity, as well as in Mg-Proto retrograde signaling.
- Chemical Biology and Proteomics Laboratory, Salk Institute for Biological Studies, La Jolla, California, USA.
Organizational Affiliation: